|
产品名称 |
AU565人乳腺癌细胞 |
|
货号 |
ZQ0379 |
|
产品介绍 |
AU565细胞系是在奥克兰海军生物科学实验室从一名乳腺癌患者的胸腔积液中提取的。该细胞系与SK-BR-3 (ATCC HTB-30)来自同一患者。 该患者是一名白人女性,43 岁,A+ 型血,曾接受放射、类固醇、环磷酰胺和 5-氟尿嘧啶治疗。AU565细胞系扩增并过表达HER-2/neu癌基因;它表达 HER-3、HER-4 和 p53 癌基因。用霉酚酸 (MPA)、佛波醇 12-肉豆蔻酸酯 13-乙酸酯 (PMA)、视黄酸 (RA) 或针对 HER-2/neu 蛋白胞外结构域的 TA-1 单克隆抗体处理细胞可诱导细胞分化。用 Neu 分化因子 (NDF) 处理 AU565 细胞也会诱导成熟表型,其特征在于细胞的形态外观。未经处理的 AU565 细胞具有致密的细胞核和薄的细胞质,而处理的细胞则表现出与分化表型相关的形态,例如被相当大的细胞质包围的扁平大细胞核。1995 年 4 月提交给 ATCC 的培养物被发现受到支原体污染,通过 BM Cycline 治疗 21 天治愈了后代。治疗六周后,通过 Hoechst 染色、PCR 和标准培养测试对细胞进行支原体检测。所有测试均为阴性。
该细胞可用作细胞生长模型来研究乳腺癌细胞的分化。 |
|
种属 |
人 |
|
性别/年龄 |
女/43岁 |
|
组织 |
胸部; 乳腺 |
|
疾病 |
腺癌 |
|
生物安全等级 |
BSL-1 |
|
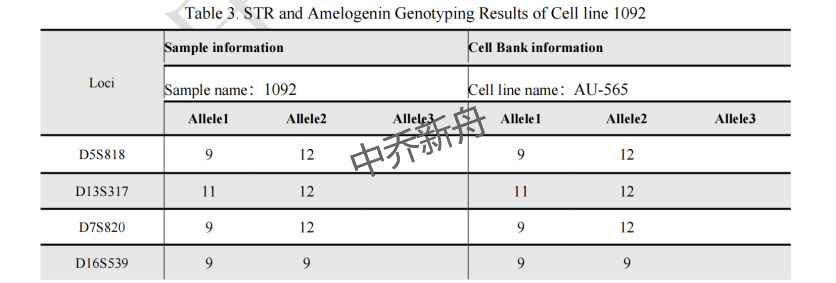
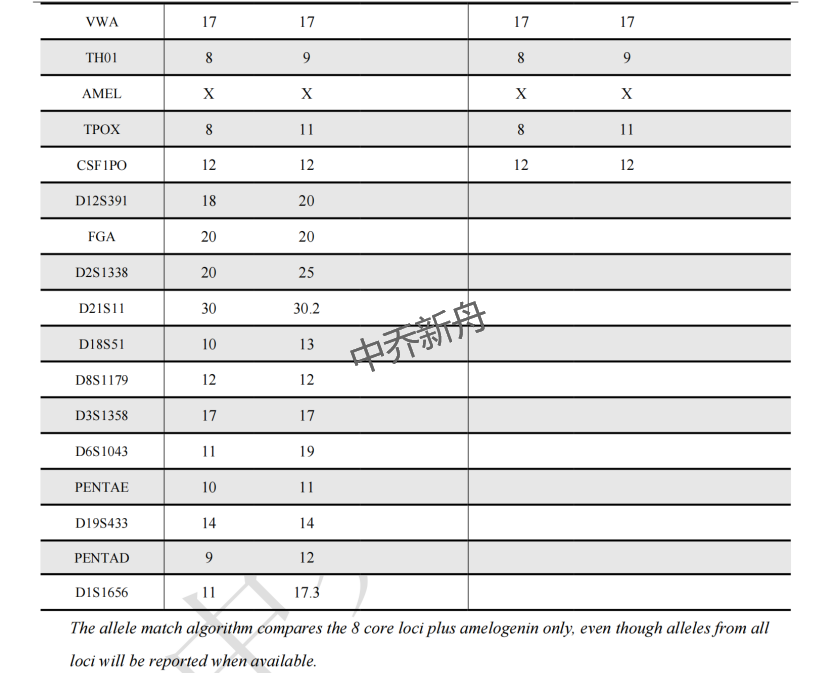
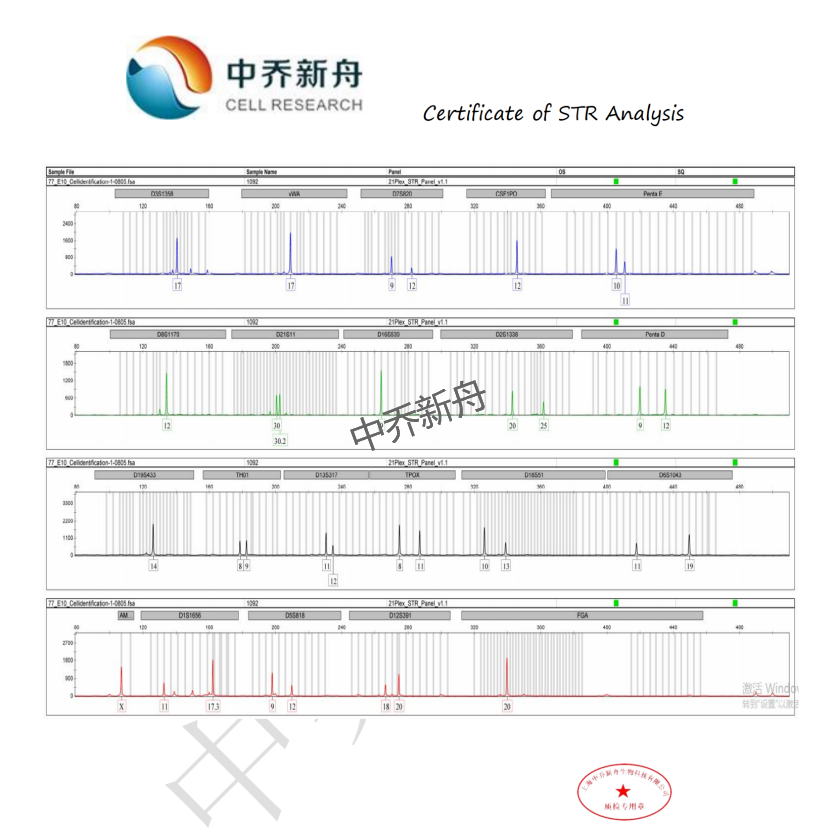
STR位点信息 |
Amelogenin: X CSF1PO: 12 D13S317: 11,12 D16S539: 9 D5S818: 9,12 D7S820: 9,12 THO1: 8,9 TPOX: 8,11 vWA: 17 |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
上皮 |
|
生长方式 |
贴壁 |
|
倍增时间 |
46.31 hours (JWGray panel) |
|
培养基和添加剂 |
RPMI-1640(中乔新舟 货号:ZQ-200)+10%胎牛血清(中乔新舟 货号:ZQ0500)+1%双抗(品牌:中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
|
|
培养条件 |
5%二氧化碳;37℃ |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ATCC;CRL-2351 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZQ0379 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial)二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司