|
产品名称 |
TOV21G人卵巢癌细胞 |
|
货号 |
ZQ1008 |
|
产品介绍 |
TOV21G是一种人卵巢癌细胞系,它源自一位62岁法裔加拿大女性的原发性恶性腺癌,于 1991 年 10 月建系。该患者没有卵巢癌的家族史。这种细胞系在生物医学研究中被广泛使用,尤其是在研究卵巢癌的生物学特性、病理机制以及药物筛选等方面。该系在染色体 3p24 处未显示缺失。 |
|
种属 |
人 |
|
性别/年龄 |
女/62岁 |
|
组织 |
卵巢 |
|
疾病 |
腺癌 |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
上皮样 |
|
生长方式 |
贴壁 |
|
倍增时间 |
1.5 days (PubMed=10949993); 27 hours (PubMed=25984343); 30.62 hours (JWGray panel) |
|
培养基和添加剂 |
MCDB105+Medium199(1:1)+15%FBS(货号:ZQ0500)+1%PS(中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
|
|
生物安全等级 |
BSL-1 |
|
培养条件 |
95%空气,5%二氧化碳;37℃ |
|
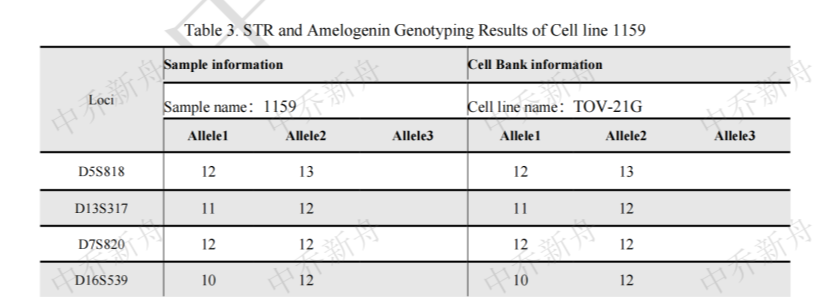
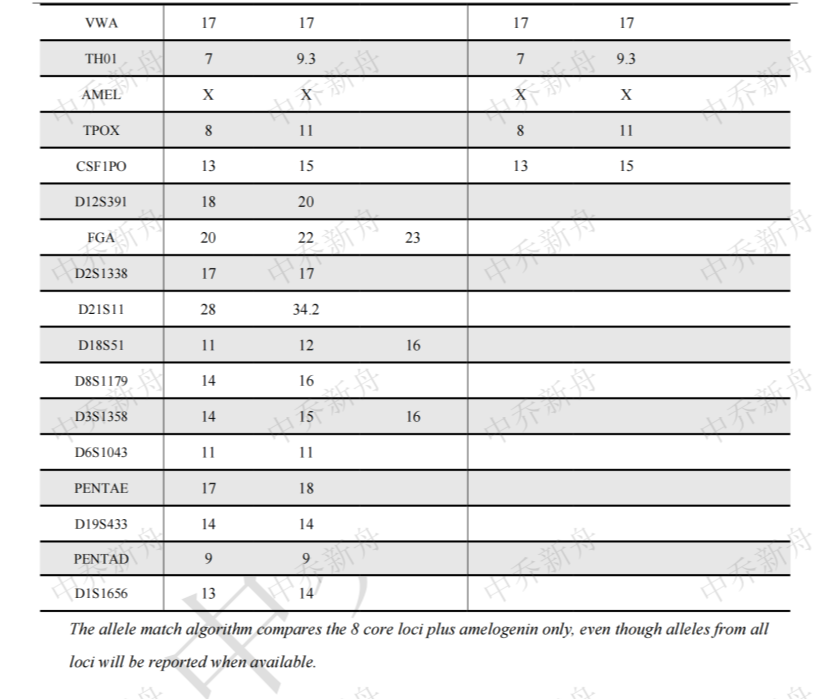
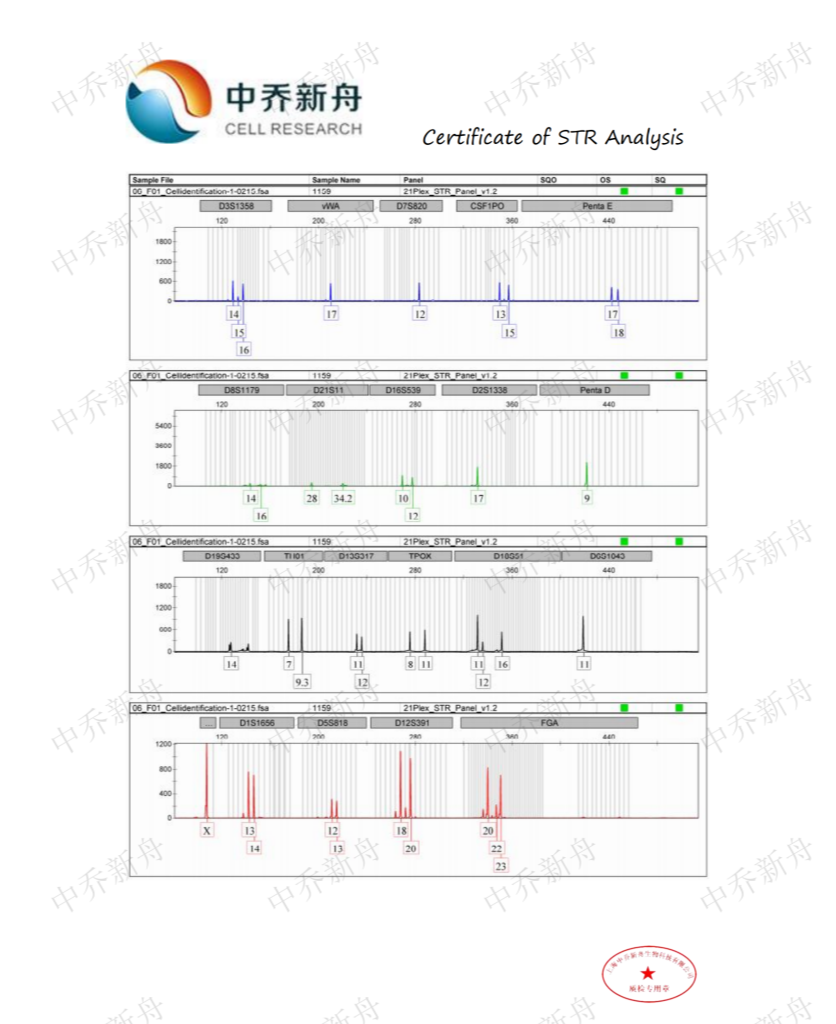
STR位点信息 |
Amelogenin :X CSF1PO: 13,15 D2S1338: 17 D3S1358: 14,16 D5S818: 12,13 D7S820 :12 D8S1179 :13,16 D13S317 :11,12 D16S539: 10,12 D18S51: 12 (PubMed=25230021; PubMed=25877200) :12,16 (PubMed=22710073) D19S433: 14,16.2 D21S11: 28,34.2 FGA: 20,23 Penta D :9 Penta E: 17,18 (PubMed=25877200) :17,19 (PubMed=25230021) TH01: 7,9.3 TPOX :8,11 vWA :17 (ATCC=CRL-3577; Cosmic-CLP=1240222; PubMed=22710073; PubMed=25230021; PubMed=25877200; PubMed=30485824) :17,18 (Direct_author_submission) |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ATCC; CRL-11730;BCRC; 60407 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZQ1008 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial )二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司