Cell culture studies provide a valuable complement to in vivo experiments, allowing for a more controlled manipulation of cellular functions and processes. For decades, cell lines have played a critical role in scientific advancements, yet researchers have become increasingly cautious when interpreting data generated from cell lines only. Factors such as misidentified and contaminated cell lines have spurred renewed interest in primary cells.1,2
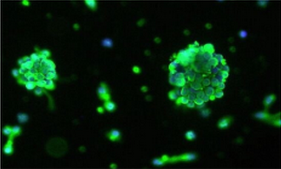
Immunofluorescence labels the glial progenitor marker A2B5 (green) and nuclei (blue) in oligospheres formed by normal primary human oligodendrocyte precursor cells. (Image by James Shen, ScienCell Research Laboratories.)
Another disadvantage of cell lines is that they often differ genetically and phenotypically from their tissue of origin. In contrast, primary cells maintain many of the important markers and functions seen in vivo.3,4Endothelial cell lines, for example, lack various functional markers, while primary endothelial cells retain these critical features.
Growing Primary Cells
Although primary cells offer many advantages, obtaining a pure population of primary cells can be a difficult and arduous process. Primary cells, in contrast to cell lines, are extremely sensitive, requiring additional nutrients not included in classical media. To optimize survival and growth, primary cells perform best in specialty media customized for each cell type. An endothelial cell, for example, has very different nutritional requirements than an epithelial cell or a neuron, and thus requires a unique medium.
Traditional cell culture media have relied on serum to provide the growth factors, hormones, lipids, and other, undefined components to support cellular growth. For primary cells, however, high serum levels can lead to differentiation or promote growth of contaminating cells like fibroblasts. In addition, the use of serum is plagued by rising costs and lot-to-lot variability. The use of specially formulated media with little or no serum circumvents these issues while enabling greater customization to promote growth of individual primary cell types.
Other practices, such as seeding primary cells on more physiologically relevant substrates rather than on synthetic polymers, can significantly improve cell attachment, growth, and purity.
Gene Expression Analysis
Gene expression analysis is critical for understanding the transcriptome profiles of primary cells and how they directly influence the cells’ functionality. Traditional reporter-gene assays and cDNA microarrays often require either transfection of exogenous material or large quantities of high-quality RNA. Primary cells, however, are notoriously difficult to transfect, and the efficiency varies greatly among cell types. In addition, primary cells have a finite lifespan and limited expansion capacity, making it difficult to obtain a high yield of RNA.
These problems can be overcome with the use of quantitative PCR (qPCR) arrays that have been validated and optimized using primary cell cDNA. qPCR arrays enable researchers to directly measure protein expression patterns of primary cells, stem cells, tissues, and cell lines. Due to the high sensitivity and specificity of these assays, even genes with very low abundance can be analyzed.
Gene expression analysis of primary cells will enable researchers to better understand biological pathways and disease processes, such as cell cycle regulation, stem cell biology, cancer development, and neurological disorders. Primary cells are critical tools in the pursuit of future scientific breakthroughs and therapies.
— Jennifer Welser-Alves, ScienCell Research Laboratories
Note
The author is Associate Director, Scientific Affairs, at ScienCell Research Laboratories, which specializes in primary cell culture. Its products include primary cells, customized culture media, and GeneQueryTM qPCR array kits.
References
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司