配套完培,省时省力,单买细胞无优惠
|
产品名称 |
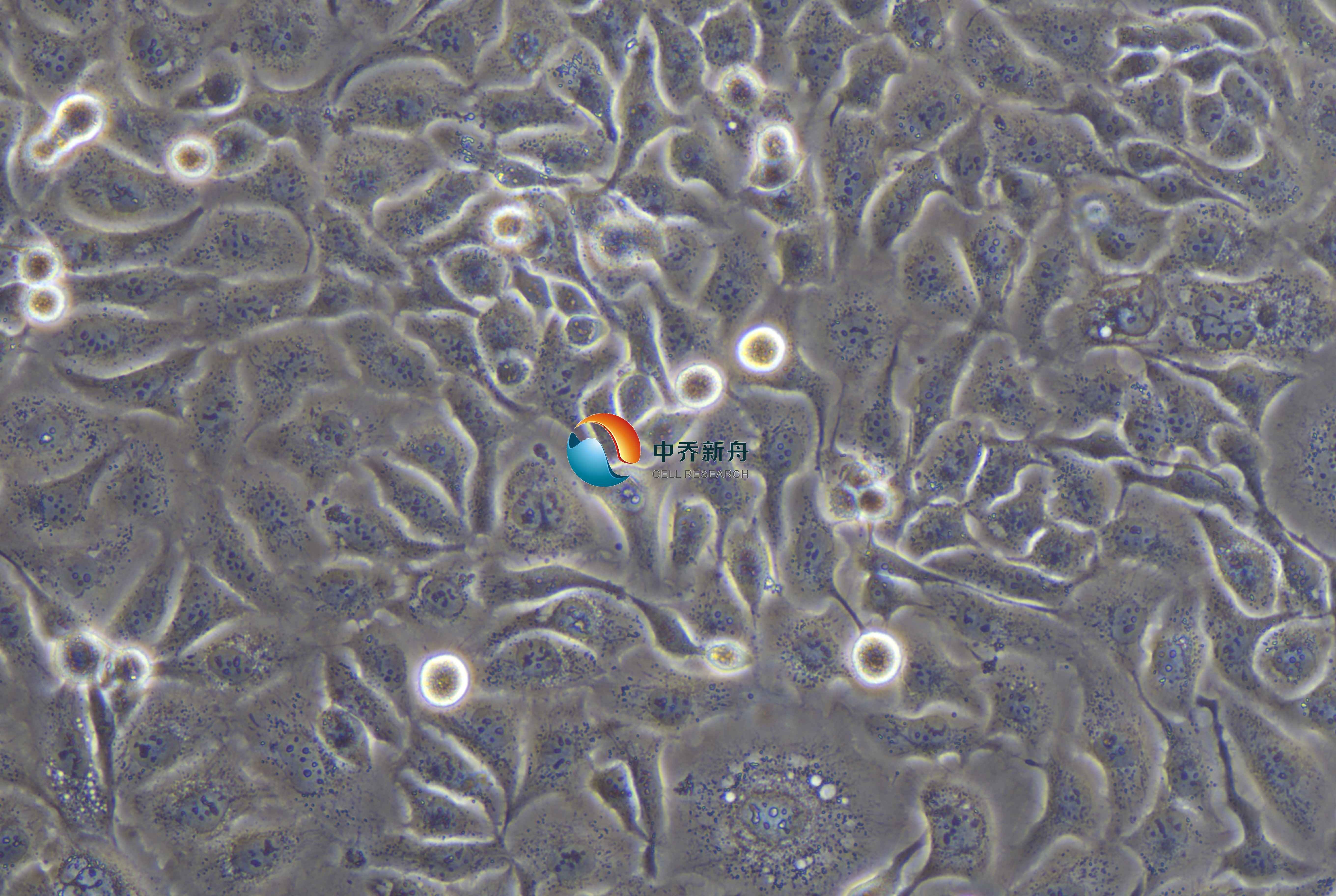
Hcc1806人乳腺癌细胞 |
|
货号 |
ZQ0769 |
|
产品介绍 |
HCC1806是一种人乳腺癌细胞系,源自一位60岁黑人女性的乳腺鳞状细胞癌(ASCC),TNM分期为IIB期,分级为2级。该细胞系于1995年7月31日建立,历时10个月。该细胞缺乏雌激素和黄体酮受体,并且缺乏表皮生长因子受体(EGFR)扩增,将其归类为三阴性乳腺癌。该细胞系有助于治疗靶点的生物学验证,因为它密切反映了TNBC在体内的行为,包括自发转移的倾向和对紫杉醇等传统疗法的耐药性。此外,干预的分子效应如AEB071治疗,对HCC1806细胞,提供了细胞增殖途径和蛋白激酶抑制剂作为治疗药物的潜力。在异种移植物模型中使用HCC1806有助于在受控环境下研究肿瘤的生长和转移。 |
|
种属 |
人 |
|
性别/年龄 |
女/60岁 |
|
组织 |
乳房;乳腺 |
|
疾病 |
棘层溶解性鳞状细胞癌;TNM舞台IIB,二级 |
|
细胞类型 |
上皮细胞 |
|
形态学 |
上皮的 |
|
生长方式 |
贴壁、附着的中等大小上皮细胞,无漂浮细胞 |
|
倍增时间 |
大约36.66小时 |
|
培养基和添加剂 |
RPMI-1640(品牌:中乔新舟 货号:ZQ-200)+10%胎牛血清(中乔新舟 货号:ZQ0500)+1%P/S(中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
|
|
生物安全等级 |
BSL-1 |
|
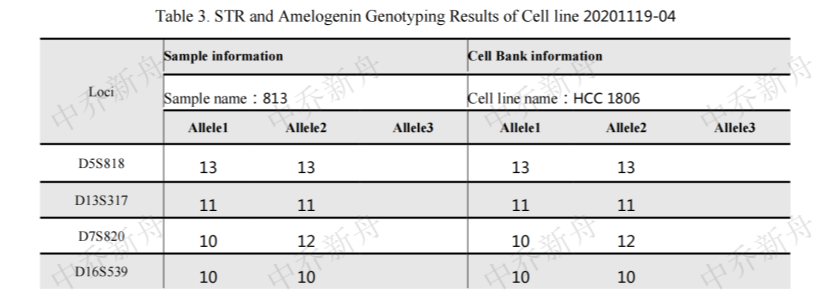
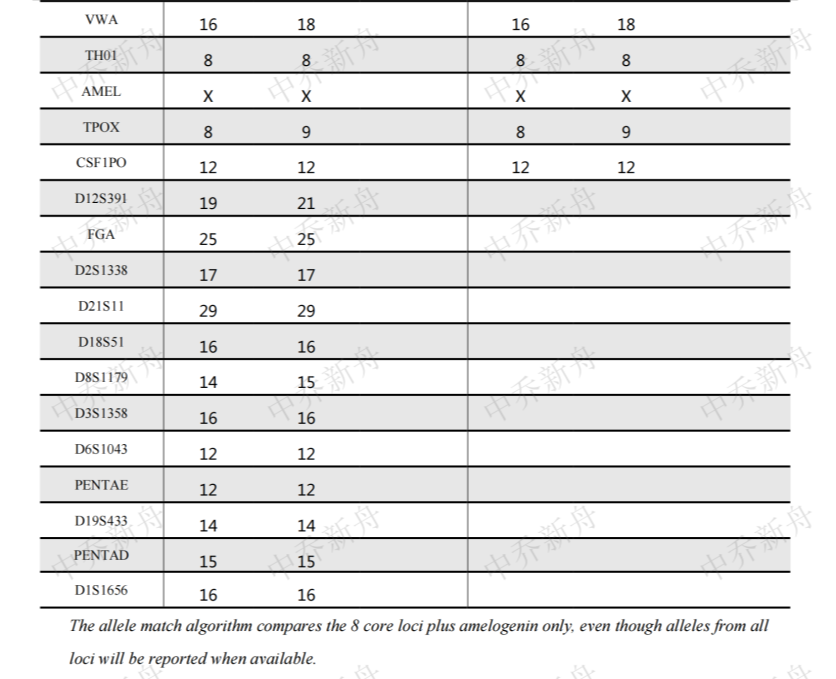
STR位点信息 |
D5S818: 13Amelogenin:X |
|
培养条件 |
95%空气,5%二氧化碳;37℃ |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ATCC; CRL-2335 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZQ0769 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial)二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |
PubMed=9288768
Ahmadian M., Wistuba I.I., Fong K.M., Behrens C., Kodagoda D.R., Saboorian M.H., Shay J.W., Tomlinson G.E., Blum J., Minna J.D., Gazdar A.F.
Analysis of the FHIT gene and FRA3B region in sporadic breast cancer, preneoplastic lesions, and familial breast cancer probands.
Cancer Res. 57:3664-3668(1997)
PubMed=9833771; DOI=10.1002/(SICI)1097-0215(19981209)78:6<766::AID-IJC15>3.0.CO;2-L
Gazdar A.F., Kurvari V., Virmani A.K., Gollahon L., Sakaguchi M., Westerfield M., Kodagoda D.R., Stasny V., Cunningham H.T., Wistuba I.I., Tomlinson G.E., Tonk V., Ashfaq R., Leitch A.M., Minna J.D., Shay J.W.
Characterization of paired tumor and non-tumor cell lines established from patients with breast cancer.
Int. J. Cancer 78:766-774(1998)
PubMed=9865903
Wistuba I.I., Behrens C., Milchgrub S., Syed S., Ahmadian M., Virmani A.K., Kurvari V., Cunningham T.H., Ashfaq R., Minna J.D., Gazdar A.F.
Comparison of features of human breast cancer cell lines and their corresponding tumors.
Clin. Cancer Res. 4:2931-2938(1998)
PubMed=11314036; DOI=10.1038/sj.onc.1204211
Forgacs E., Wren J.D., Kamibayashi C., Kondo M., Xu X.L., Markowitz S.D., Tomlinson G.E., Muller C.Y., Gazdar A.F., Garner H.R., Minna J.D.
Searching for microsatellite mutations in coding regions in lung, breast, ovarian and colorectal cancers.
Oncogene 20:1005-1009(2001)
PubMed=12353263; DOI=10.1002/gcc.10107
Popovici C., Basset C., Bertucci F., Orsetti B., Adelaide J., Mozziconacci M.-J., Conte N., Murati A., Ginestier C., Charafe-Jauffret E., Ethier S.P., Lafage-Pochitaloff M., Theillet C., Birnbaum D., Chaffanet M.
Reciprocal translocations in breast tumor cell lines: cloning of a t(3;20) that targets the FHIT gene.
Genes Chromosomes Cancer 35:204-218(2002)
PubMed=12800145; DOI=10.1002/gcc.10218
Adelaide J., Huang H.-E., Murati A., Alsop A.E., Orsetti B., Mozziconacci M.-J., Popovici C., Ginestier C., Letessier A., Basset C., Courtay-Cahen C., Jacquemier J., Theillet C., Birnbaum D., Edwards P.A.W., Chaffanet M.
A recurrent chromosome translocation breakpoint in breast and pancreatic cancer cell lines targets the neuregulin/NRG1 gene.
Genes Chromosomes Cancer 37:333-345(2003)
PubMed=19582160; DOI=10.1371/journal.pone.0006146
Kao J., Salari K., Bocanegra M., Choi Y.-L., Girard L., Gandhi J., Kwei K.A., Hernandez-Boussard T., Wang P., Gazdar A.F., Minna J.D., Pollack J.R.
Molecular profiling of breast cancer cell lines defines relevant tumor models and provides a resource for cancer gene discovery.
PLoS ONE 4:E6146-E6146(2009)
PubMed=20070913; DOI=10.1186/1471-2407-10-15
Tsuji K., Kawauchi S., Saito S., Furuya T., Ikemoto K., Nakao M., Yamamoto S., Oka M., Hirano T., Sasaki K.
Breast cancer cell lines carry cell line-specific genomic alterations that are distinct from aberrations in breast cancer tissues: comparison of the CGH profiles between cancer cell lines and primary cancer tissues.
BMC Cancer 10:15.1-15.10(2010)
PubMed=20164919; DOI=10.1038/nature08768
Bignell G.R., Greenman C.D., Davies H., Butler A.P., Edkins S., Andrews J.M., Buck G., Chen L., Beare D., Latimer C., Widaa S., Hinton J., Fahey C., Fu B.-Y., Swamy S., Dalgliesh G.L., Teh B.T., Deloukas P., Yang F.-T., Campbell P.J., Futreal P.A., Stratton M.R.
Signatures of mutation and selection in the cancer genome.
Nature 463:893-898(2010)
PubMed=20679594; DOI=10.1093/jnci/djq279
Gazdar A.F., Girard L., Lockwood W.W., Lam W.L., Minna J.D.
Lung cancer cell lines as tools for biomedical discovery and research.
J. Natl. Cancer Inst. 102:1310-1321(2010)
PubMed=21778573; DOI=10.3233/BD-2010-0307
Chavez K.J., Garimella S.V., Lipkowitz S.
Triple negative breast cancer cell lines: one tool in the search for better treatment of triple negative breast cancer.
Breast Dis. 32:35-48(2010)
PubMed=21603256; DOI=10.4137/BCBCR.S7087
Neves L.A.H., Ingram L.M., Davis M.B.
The characterization of cell line CRL-2335 as a basal-like breast carcinoma model.
Breast Cancer (Auckl.) 5:67-72(2011)
PubMed=22460905; DOI=10.1038/nature11003
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=22585861; DOI=10.1158/2159-8290.CD-11-0224
Marcotte R., Brown K.R., Suarez Saiz F.J., Sayad A., Karamboulas K., Krzyzanowski P.M., Sircoulomb F., Medrano M., Fedyshyn Y., Koh J.L.-Y., van Dyk D., Fedyshyn B., Luhova M., Brito G.C., Vizeacoumar F.J., Vizeacoumar F.S., Datti A., Kasimer D., Buzina A., Mero P., Misquitta C., Normand J., Haider M., Ketela T., Wrana J.L., Rottapel R., Neel B.G., Moffat J.
Essential gene profiles in breast, pancreatic, and ovarian cancer cells.
Cancer Discov. 2:172-189(2012)
PubMed=23601657; DOI=10.1186/bcr3415
Riaz M., van Jaarsveld M.T.M., Hollestelle A., Prager-van der Smissen W.J.C., Heine A.A.J., Boersma A.W.M., Liu J.-J., Helmijr J.C.A., Ozturk B., Smid M., Wiemer E.A.C., Foekens J.A., Martens J.W.M.
miRNA expression profiling of 51 human breast cancer cell lines reveals subtype and driver mutation-specific miRNAs.
Breast Cancer Res. 15:R33.1-R33.17(2013)
PubMed=24176112; DOI=10.1186/gb-2013-14-10-r110
Daemen A., Griffith O.L., Heiser L.M., Wang N.J., Enache O.M., Sanborn Z., Pepin F., Durinck S., Korkola J.E., Griffith M., Hur J.S., Huh N., Chung J., Cope L., Fackler M.J., Umbricht C.B., Sukumar S., Seth P., Sukhatme V.P., Jakkula L.R., Lu Y.-L., Mills G.B., Cho R.J., Collisson E.A., van 't Veer L.J., Spellman P.T., Gray J.W.
Modeling precision treatment of breast cancer.
Genome Biol. 14:R110.1-R110.14(2013)

 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司