|
产品名称 |
SU-DHL-4人B细胞淋巴瘤 |
|
货号 |
ZQ0982 |
|
产品介绍 |
该细胞系建系于1976年,来自38岁白人男性的腹膜渗出物。这个细胞系代表了弥漫性大b细胞淋巴瘤(DLBCL)的模型,DLBCL是成人中最常见的非霍奇金淋巴瘤之一。该细胞系的建立为DLBCL的生物学,特别是淋巴瘤发生和肿瘤进展的细胞和分子机制提供了有价值的见解。该细胞系有14号、18号染色体易位。在BCL-2基因中有一个主要的重排。该细胞系对念珠菌摄入呈阴性。有资料显示该细胞对爱泼斯坦-巴尔病毒(EBV)呈阴性。每个批次均通过本库支原体检测,结果阴性。
在研究中,SU-DHL-4细胞被广泛用于研究各种化疗药物和靶向治疗药物的疗效和作用机制,反映了其在淋巴瘤治疗研究中的重要性。这些细胞表达一些与b细胞谱系相关的关键免疫表型标记,如CD19和CD20,它们对b淋巴细胞的发育和功能至关重要。这些标记也使SU-DHL-4成为测试b细胞特异性疗法的极好靶点,包括单克隆抗体和小分子抑制剂,它们破坏淋巴瘤细胞存活和增殖的关键信号通路。 注意事项:建议接种密度为10万至20万/ml,建议传代密度为80万 至100万/ml。 |
|
种属 |
人 |
|
性别/年龄 |
男性/38岁 |
|
组织 |
腹腔积液 |
|
疾病 |
弥漫性大B细胞淋巴瘤 |
|
生物安全等级 |
BSL-1 |
|
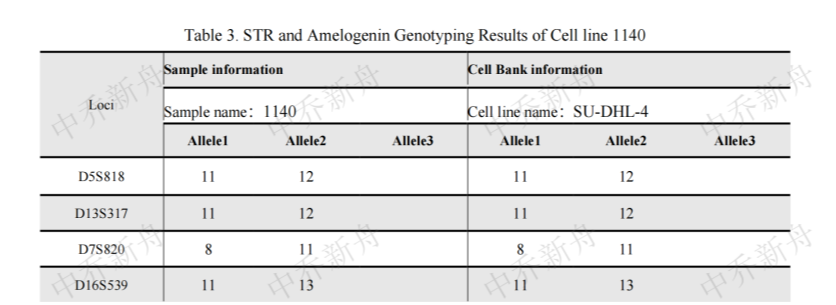
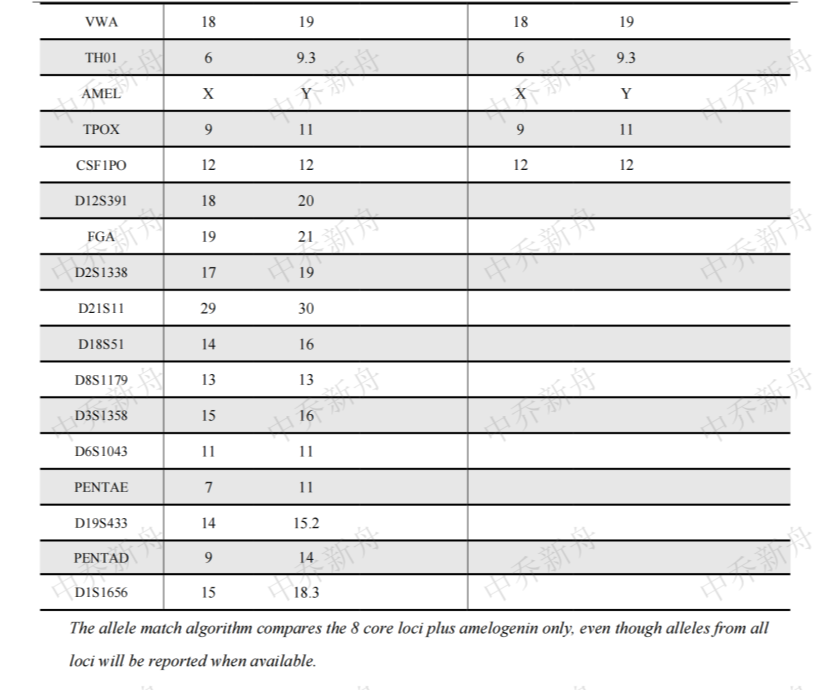
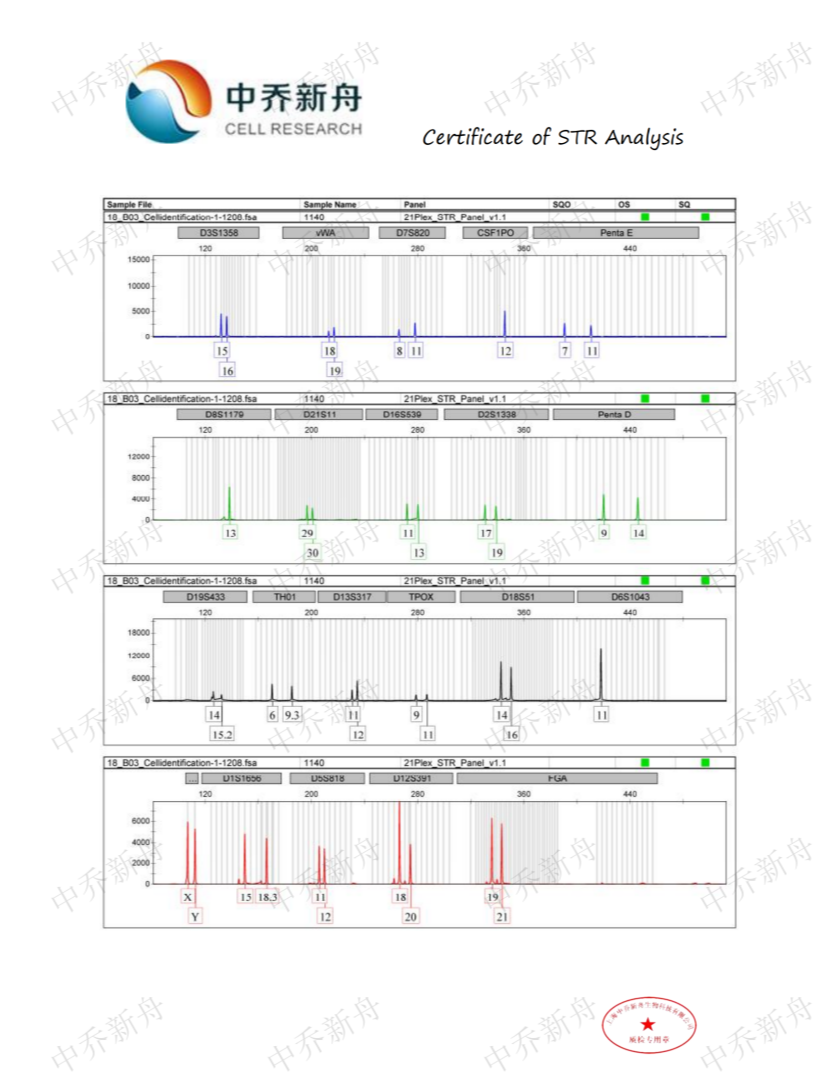
STR位点信息 |
D5S818: 11, 12 |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
淋巴母细胞样 |
|
生长方式 |
悬浮 |
|
倍增时间 |
~40 hours (DSMZ=ACC-495) |
|
培养基和添加剂 |
RPMI-1640(中乔新舟 货号:ZQ-200)+10%FBS(品牌:中乔新舟 货号:ZQ0500)+1%PS(中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
|
|
培养条件 |
95%空气,5%二氧化碳;37℃ |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ATCC; CRL-2957 DSMZ; ACC-495 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZQ0982 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial )二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |
PubMed=177185; DOI=10.1002/1097-0142(197605)37:5<2158::AID-CNCR2820370503>3.0.CO;2-F
Epstein A.L., Herman M.M., Kim H., Dorfman R.F., Kaplan H.S.
Biology of the human malignant lymphomas. III. Intracranial heterotransplantation in the nude, athymic mouse.
Cancer 37:2158-2176(1976)
PubMed=214220; DOI=10.1002/1097-0142(197811)42:5<2379::AID-CNCR2820420539>3.0.CO;2-4
Epstein A.L., Levy R., Kim H., Henle W., Henle G., Kaplan H.S.
Biology of the human malignant lymphomas. IV. Functional characterization of ten diffuse histiocytic lymphoma cell lines.
Cancer 42:2379-2391(1978)
PubMed=83902; DOI=10.1002/1097-0142(197901)43:1<1::AID-CNCR2820430102>3.0.CO;2-M
Kaplan H.S., Goodenow R.S., Gartner S., Bieber M.M.
Biology and virology of the human malignant lymphomas: 1st Milford D Schulz Lecture.
Cancer 43:1-24(1979)
PubMed=371794
Epstein A.L., Kaplan H.S.
Feeder layer and nutritional requirements for the establishment and cloning of human malignant lymphoma cell lines.
Cancer Res. 39:1748-1759(1979)
PubMed=2985879; DOI=10.1016/0145-2126(85)90084-0
Drexler H.G., Gaedicke G., Minowada J.
Isoenzyme studies in human leukemia-lymphoma cell lines -- 1 carboxylic esterase.
Leuk. Res. 9:209-229(1985)
PubMed=3159941; DOI=10.1016/0145-2126(85)90134-1
Drexler H.G., Gaedicke G., Minowada J.
Isoenzyme studies in human leukemia-lymphoma cell lines -- III Beta-hexosaminidase (E.C. 3.2.1.30).
Leuk. Res. 9:549-559(1985)
PubMed=3874327; DOI=10.1016/0145-2126(85)90133-x
Drexler H.G., Gaedicke G., Minowada J.
Isoenzyme studies in human leukemia-lymphoma cells lines -- II. Acid phosphatase.
Leuk. Res. 9:537-548(1985)
PubMed=3881165; DOI=10.1016/0165-4608(85)90186-4
Kaiser-McCaw Hecht B., Epstein A.L., Berger C.S., Kaplan H.S., Hecht F.
Histiocytic lymphoma cell lines: immunologic and cytogenetic studies.
Cancer Genet. Cytogenet. 14:205-218(1985)
PubMed=2875799; DOI=10.1016/0092-8674(86)90362-4
Cleary M.L., Smith S.D., Sklar J.
Cloning and structural analysis of cDNAs for bcl-2 and a hybrid bcl-2/immunoglobulin transcript resulting from the t(14;18) translocation.
Cell 47:19-28(1986)
PubMed=8547074; DOI=10.1111/j.1365-2141.1995.tb05302.x
Siebert R., Willers C.P., Schramm A., Fossa A., Dresen I.M.G., Uppenkamp M., Nowrousian M.R., Seeber S., Opalka B.
Homozygous loss of the MTS1/p16 and MTS2/p15 genes in lymphoma and lymphoblastic leukaemia cell lines.
Br. J. Haematol. 91:350-354(1995)
DOI=10.11418/jtca1981.15.4_211
Matsuo Y., Okochi A., Ariyasu T., Iimura E., Ohno T.
Identification of cell lines with variable numbers of tandem repeat (VNTR) amplified by polymerase chain reaction.
Tissue Cult. Res. Commun. 15:211-219(1996)
PubMed=8957066; DOI=10.1111/j.1349-7006.1996.tb03112.x
Kawasaki N., Matsuo Y., Yoshino T., Yanai H., Oka T., Teramoto N., Liu C., Kondo E., Minowada J., Akagi T.
Metastatic potential of lymphoma/leukemia cell lines in SCID mice is closely related to expression of CD44.
Jpn. J. Cancer Res. 87:1070-1077(1996)
PubMed=9738977; DOI=10.1111/j.1349-7006.1998.tb03275.x
Takizawa J., Suzuki R., Kuroda H., Utsunomiya A., Kagami Y., Joh T., Aizawa Y., Ueda R., Seto M.
Expression of the TCL1 gene at 14q32 in B-cell malignancies but not in adult T-cell leukemia.
Jpn. J. Cancer Res. 89:712-718(1998)
PubMed=9787181; DOI=10.1182/blood.V92.9.3410
Sakai A., Thieblemont C., Wellmann A., Jaffe E.S., Raffeld M.
PTEN gene alterations in lymphoid neoplasms.
Blood 92:3410-3415(1998)
PubMed=10739008; DOI=10.1016/S0145-2126(99)00182-4
Inoue K., Kohno T., Takakura S., Hayashi Y., Mizoguchi H., Yokota J.
Frequent microsatellite instability and BAX mutations in T cell acute lymphoblastic leukemia cell lines.
Leuk. Res. 24:255-262(2000)
DOI=10.1016/B978-0-12-221970-2.50457-5
Drexler H.G.
The leukemia-lymphoma cell line factsbook.
(In) ISBN 9780122219702; pp.1-733; Academic Press; London (2001)
PubMed=11226526; DOI=10.1016/S0145-2126(00)00121-1
Inoue K., Kohno T., Takakura S., Hayashi Y., Mizoguchi H., Yokota J.
Corrigendum to: Frequent microsatellite instability and BAX mutations in T cell acute lymphoblastic leukemia cell lines Leukemia Research 24 (2000),255-262.
Leuk. Res. 25:275-278(2001)
PubMed=12169673; DOI=10.1016/S1525-1578(10)60693-9
Robetorye R.S., Bohling S.D., Morgan J.W., Fillmore G.C., Lim M.S., Elenitoba-Johnson K.S.J.
Microarray analysis of B-cell lymphoma cell lines with the t(14;18).
J. Mol. Diagn. 4:123-136(2002)
PubMed=17363600; DOI=10.1158/0008-5472.CAN-06-3254
Strauss S.J., Higginbottom K., Juliger S., Maharaj L., Allen P., Schenkein D., Lister T.A., Joel S.P.
The proteasome inhibitor bortezomib acts independently of p53 and induces cell death via apoptosis and mitotic catastrophe in B-cell lymphoma cell lines.
Cancer Res. 67:2783-2790(2007)
PubMed=18357372; DOI=10.3892/or.19.4.889
Pop I., Pop L., Vitetta E.S., Ghetie M.-A.
Generation of multidrug resistant lymphoma cell lines stably expressing P-glycoprotein.
Oncol. Rep. 19:889-895(2008)
PubMed=19278952; DOI=10.1182/blood-2009-01-202028
Li C., Kim S.-W., Rai D., Bolla A.R., Adhvaryu S., Kinney M.C., Robetorye R.S., Aguiar R.C.T.
Copy number abnormalities, MYC activity, and the genetic fingerprint of normal B cells mechanistically define the microRNA profile of diffuse large B-cell lymphoma.
Blood 113:6681-6690(2009)
PubMed=20054396; DOI=10.1038/nature08638
Davis R.E., Ngo V.N., Lenz G., Tolar P., Young R.M., Romesser P.B., Kohlhammer H., Lamy L., Zhao H., Yang Y.-D., Xu W.-H., Shaffer A.L. III, Wright G., Xiao W.-M., Powell J.I., Jiang J.-K., Thomas C.J., Rosenwald A., Ott G., Muller-Hermelink H.-K., Gascoyne R.D., Connors J.M., Johnson N.A., Rimsza L.M., Campo E., Jaffe E.S., Wilson W.H., Delabie J., Smeland E.B., Fisher R.I., Braziel R.M., Tubbs R.R., Cook J.R., Weisenburger D.D., Chan W.C., Pierce S.K., Staudt L.M.
Chronic active B-cell-receptor signalling in diffuse large B-cell lymphoma.
Nature 463:88-92(2010)
PubMed=20628145; DOI=10.1182/blood-2010-05-282780
Green M.R., Monti S., Rodig S.J., Juszczynski P., Currie T., O'Donnell E., Chapuy B., Takeyama K., Neuberg D., Golub T.R., Kutok J.L., Shipp M.A.
Integrative analysis reveals selective 9p24.1 amplification, increased PD-1 ligand expression, and further induction via JAK2 in nodular sclerosing Hodgkin lymphoma and primary mediastinal large B-cell lymphoma.
Blood 116:3268-3277(2010)
PubMed=20889926; DOI=10.1182/blood-2010-06-290437
Pham L.V., Fu L.-C., Tamayo A.T., Bueso-Ramos C.E., Drakos E., Vega-Vazquez F., Medeiros L.J., Ford R.J. Jr.
Constitutive BR3 receptor signaling in diffuse, large B-cell lymphomas stabilizes nuclear factor-kappaB-inducing kinase while activating both canonical and alternative nuclear factor-kappaB pathways.
Blood 117:200-210(2011)
PubMed=22460905; DOI=10.1038/nature11003
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=23292937; DOI=10.1073/pnas.1205299110
Zhang J., Grubor V., Love C.L., Banerjee A., Richards K.L., Mieczkowski P.A., Dunphy C., Choi W., Au W.Y., Srivastava G., Lugar P.L., Rizzieri D.A., Lagoo A.S., Bernal-Mizrachi L., Mann K.P., Flowers C.R., Naresh K., Evens A.M., Gordon L.I., Czader M.B., Gill J.I., Hsi E.D., Liu Q.-Q., Fan A., Walsh K., Jima D., Smith L.L., Johnson A.J., Byrd J.C., Luftig M.A., Ni T., Zhu J., Chadburn A., Levy S., Dunson D.B., Dave S.S.
Genetic heterogeneity of diffuse large B-cell lymphoma.
Proc. Natl. Acad. Sci. U.S.A. 110:1398-1403(2013)
PubMed=25355872; DOI=10.1128/JVI.02570-14
Cao S.-B., Strong M.J., Wang X., Moss W.N., Concha M., Lin Z., O'Grady T., Baddoo M., Fewell C., Renne R., Flemington E.K.
High-throughput RNA sequencing-based virome analysis of 50 lymphoma cell lines from the Cancer Cell Line Encyclopedia project.
J. Virol. 89:713-729(2015)
PubMed=25485619; DOI=10.1038/nbt.3080
Klijn C., Durinck S., Stawiski E.W., Haverty P.M., Jiang Z.-S., Liu H.-B., Degenhardt J., Mayba O., Gnad F., Liu J.-F., Pau G., Reeder J., Cao Y., Mukhyala K., Selvaraj S.K., Yu M.-M., Zynda G.J., Brauer M.J., Wu T.D., Gentleman R.C., Manning G., Yauch R.L., Bourgon R., Stokoe D., Modrusan Z., Neve R.M., de Sauvage F.J., Settleman J., Seshagiri S., Zhang Z.-M.
A comprehensive transcriptional portrait of human cancer cell lines.
Nat. Biotechnol. 33:306-312(2015)
PubMed=25877200; DOI=10.1038/nature14397
Yu M., Selvaraj S.K., Liang-Chu M.M.Y., Aghajani S., Busse M., Yuan J., Lee G., Peale F.V., Klijn C., Bourgon R., Kaminker J.S., Neve R.M.
A resource for cell line authentication, annotation and quality control.
Nature 520:307-311(2015)
PubMed=25894527; DOI=10.1371/journal.pone.0121314
Bausch-Fluck D., Hofmann A., Bock T., Frei A.P., Cerciello F., Jacobs A., Moest H., Omasits U., Gundry R.L., Yoon C., Schiess R., Schmidt A., Mirkowska P., Hartlova A.S., Van Eyk J.E., Bourquin J.-P., Aebersold R., Boheler K.R., Zandstra P.W., Wollscheid B.
A mass spectrometric-derived cell surface protein atlas.
PLoS ONE 10:E0121314-E0121314(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=26727417; DOI=10.3109/10428194.2015.1108414
Drexler H.G., Eberth S., Nagel S., MacLeod R.A.F.
Malignant hematopoietic cell lines: in vitro models for double-hit B-cell lymphomas.
Leuk. Lymphoma 57:1015-1020(2016)
PubMed=27397505; DOI=10.1016/j.cell.2016.06.017
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., Aben N., Goncalves E., Barthorpe S., Lightfoot H., Cokelaer T., Greninger P., van Dyk E., Chang H., de Silva H., Heyn H., Deng X.-M., Egan R.K., Liu Q.-S., Mironenko T., Mitropoulos X., Richardson L., Wang J.-H., Zhang T.-H., Moran S., Sayols S., Soleimani M., Tamborero D., Lopez-Bigas N., Ross-Macdonald P., Esteller M., Gray N.S., Haber D.A., Stratton M.R., Benes C.H., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Garnett M.J.
A landscape of pharmacogenomic interactions in cancer.
Cell 166:740-754(2016)
PubMed=28196595; DOI=10.1016/j.ccell.2017.01.005
Li J., Zhao W., Akbani R., Liu W.-B., Ju Z.-L., Ling S.-Y., Vellano C.P., Roebuck P., Yu Q.-H., Eterovic A.K., Byers L.A., Davies M.A., Deng W.-L., Gopal Y.N.V., Chen G., von Euw E.M., Slamon D.J., Conklin D., Heymach J.V., Gazdar A.F., Minna J.D., Myers J.N., Lu Y.-L., Mills G.B., Liang H.
Characterization of human cancer cell lines by reverse-phase protein arrays.
Cancer Cell 31:225-239(2017)
PubMed=28356514; DOI=10.1073/pnas.1700682114
Chen J., Zhang Y., Petrus M.N., Xiao W.-M., Nicolae A., Raffeld M., Pittaluga S., Bamford R.N., Nakagawa M., Ouyang S.T.-Y., Epstein A.L., Kadin M.E., Del Mistro A., Woessner R.D., Jaffe E.S., Waldmann T.A.
Cytokine receptor signaling is required for the survival of ALK- anaplastic large cell lymphoma, even in the presence of JAK1/STAT3 mutations.
Proc. Natl. Acad. Sci. U.S.A. 114:3975-3980(2017)
PubMed=29416618; DOI=10.18632/oncotarget.20378
Liu W., Chen J., Tamayo A.T., Ruan C.-G., Li L., Zhou S.-H., Shen C., Young K.H., Westin J., Davis R.E., Hu S.-M., Medeiros L.J., Ford R.J. Jr., Pham L.V.
Preclinical efficacy and biological effects of the oral proteasome inhibitor ixazomib in diffuse large B-cell lymphoma.
Oncotarget 9:346-360(2018)
PubMed=29666304; DOI=10.1158/1078-0432.CCR-17-3004
Pham L.V., Huang S.-J., Zhang H., Zhang J., Bell T., Zhou S.-H., Pogue E., Ding Z.-Y., Lam L., Westin J., Davis R.E., Young K.H., Medeiros L.J., Ford R.J. Jr., Nomie K., Zhang L., Wang M.
Strategic therapeutic targeting to overcome venetoclax resistance in aggressive B-cell lymphomas.
Clin. Cancer Res. 24:3967-3980(2018)
PubMed=30165192; DOI=10.1016/j.canlet.2018.08.020
Qu C.-J., Kunkalla K., Vaghefi A., Frederiksen J.K., Liu Y.-D., Chapman J.R., Blonska M., Bernal-Mizrachi L., Alderuccio J.P., Lossos I.S., Landgraf R., Vega-Vazquez F.
Smoothened stabilizes and protects TRAF6 from degradation: a novel non-canonical role of smoothened with implications in lymphoma biology.
Cancer Lett. 436:149-158(2018)
PubMed=30285677; DOI=10.1186/s12885-018-4840-5
Tan K.-T., Ding L.-W., Sun Q.-Y., Lao Z.-T., Chien W., Ren X., Xiao J.-F., Loh X.-Y., Xu L., Lill M., Mayakonda A., Lin D.-C., Yang H., Koeffler H.P.
Profiling the B/T cell receptor repertoire of lymphocyte derived cell lines.
BMC Cancer 18:940.1-940.13(2018)
PubMed=30629668; DOI=10.1371/journal.pone.0210404
Uphoff C.C., Pommerenke C., Denkmann S.A., Drexler H.G.
Screening human cell lines for viral infections applying RNA-Seq data analysis.
PLoS ONE 14:E0210404-E0210404(2019)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. III, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=31160637; DOI=10.1038/s41598-019-44491-x
Quentmeier H., Pommerenke C., Dirks W.G., Eberth S., Koeppel M., MacLeod R.A.F., Nagel S., Steube K., Uphoff C.C., Drexler H.G.
The LL-100 panel: 100 cell lines for blood cancer studies.
Sci. Rep. 9:8218-8218(2019)
PubMed=31978347; DOI=10.1016/j.cell.2019.12.023
Nusinow D.P., Szpyt J., Ghandi M., Rose C.M., McDonald E.R. III, Kalocsay M., Jane-Valbuena J., Gelfand E.T., Schweppe D.K., Jedrychowski M.P., Golji J., Porter D.A., Rejtar T., Wang Y.K., Kryukov G.V., Stegmeier F., Erickson B.K., Garraway L.A., Sellers W.R., Gygi S.P.
Quantitative proteomics of the Cancer Cell Line Encyclopedia.
Cell 180:387-402.e16(2020)
PubMed=35839778; DOI=10.1016/j.ccell.2022.06.010
Goncalves E., Poulos R.C., Cai Z.-X., Barthorpe S., Manda S.S., Lucas N., Beck A., Bucio-Noble D., Dausmann M., Hall C., Hecker M., Koh J., Lightfoot H., Mahboob S., Mali I., Morris J., Richardson L., Seneviratne A.J., Shepherd R., Sykes E., Thomas F., Valentini S., Williams S.G., Wu Y.-X., Xavier D., MacKenzie K.L., Hains P.G., Tully B., Robinson P.J., Zhong Q., Garnett M.J., Reddel R.R.
Pan-cancer proteomic map of 949 human cell lines.
Cancer Cell 40:835-849.e8(2022)
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司