|
产品名称 |
T98G (人脑胶质瘤细胞) |
|
货号 |
ZQ0984 |
|
产品说明 |
T98G 人脑胶质瘤细胞是一种从人类脑肿瘤组织中分离并永生化的细胞系,用于科学研究。这种细胞系来源于一位61岁的白人男性多形性胶质母细胞瘤(glioblastoma multiforme, GBM)患者的大脑。T98G细胞具有成纤维细胞样的形态,并且在体外培养时表现出锚定独立的生长特性,即它们能够在悬浮状态下生长。当缺乏血清或拥挤时,细胞进入存活的 G1 停滞状态。 T98G细胞在多个研究领域中都有应用,包括药物筛选、基因功能研究、癌症生物学研究等。这些细胞提供了一个有用的模型系统,用于研究人类胶质瘤的生物学特性和开发新的治疗方法。 |
|
种属 |
人 |
|
性别/年龄 |
男/61岁 |
|
组织 |
脑 |
|
疾病 |
胶质母细胞瘤 |
|
生物安全等级 |
BSL-1 |
|
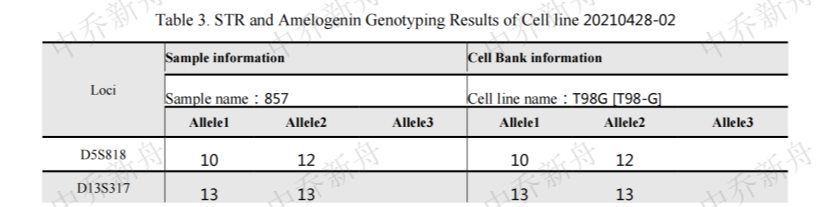
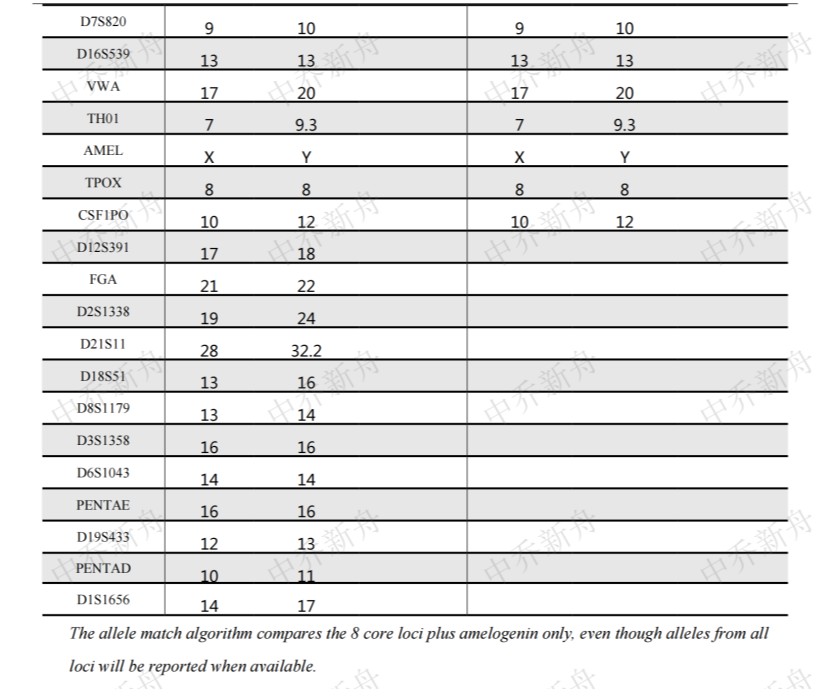
STR位点信息 |
Amelogenin: X,Y CSF1PO: 10,12 D13S317: 13 D16S539: 13 D5S818: 10,12 D7S820: 9,10 TH01: 7,9.3 TPOX: 8 vWA: 17,20 |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
成纤维细胞样 |
|
生长方式 |
贴壁细胞 |
|
倍增时间 |
22 hours (PubMed=9842975); 40 hours (PubMed=25984343); 26-36 hours (PubMed=30188626); ~28 hours (PBCF) |
|
培养基和添加剂 |
MEM(含有NEAA)基础培养基(中乔新舟 货号:ZQ-300)+10%胎牛血清(中乔新舟 货号:ZQ500-A)+1%PS(中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
|
|
培养条件 |
5%二氧化碳;37℃ |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ATCC; CRL-1690 ECACC; 92090213 JCRB; IFO50303 JCRB; JCRB9041 KCLB; 21690 RCB; RCB1954 |
|
货号 |
ZQ0984 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial )二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |
PubMed=222778; DOI=10.1002/jcp.1040990107
Stein G.H.
T98G: an anchorage-independent human tumor cell line that exhibits stationary phase G1 arrest in vitro.
J. Cell. Physiol. 99:43-54(1979)
PubMed=3413074; DOI=10.1073/pnas.85.16.6042
Pereira-Smith O.M., Smith J.R.
Genetic analysis of indefinite division in human cells: identification of four complementation groups.
Proc. Natl. Acad. Sci. U.S.A. 85:6042-6046(1988)
PubMed=9090379; DOI=10.1038/ng0497-356
Steck P.A., Pershouse M.A., Jasser S.A., Yung W.-K.A., Lin H., Ligon A.H., Langford L.A., Baumgard M.L., Hattier T., Davis T., Frye C., Hu R., Swedlund B., Teng D.H.-F., Tavtigian S.V.
Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers.
Nat. Genet. 15:356-362(1997)
PubMed=9842975; DOI=10.1002/(SICI)1097-0215(19981218)79:6<640::AID-IJC15>3.0.CO;2-z
Weller M., Rieger J., Grimmel C., Van Meir E.G., De Tribolet N., Krajewski S., Reed J.C., von Deimling A., Dichgans J.
Predicting chemoresistance in human malignant glioma cells: the role of molecular genetic analyses.
Int. J. Cancer 79:640-644(1998)
DOI=10.1007/0-306-46861-1_11
Ali-Osman F.
Brain tumors.
(In) Human cell culture. Vol. 2. Cancer cell lines part 2; Masters J.R.W., Palsson B.O. (eds.); pp.167-184; Kluwer Academic Publishers; New York (1999)
PubMed=10416987; DOI=10.1111/j.1750-3639.1999.tb00536.x
Ishii N., Maier D., Merlo A., Tada M., Sawamura Y., Diserens A.-C., Van Meir E.G.
Frequent co-alterations of TP53, p16/CDKN2A, p14ARF, PTEN tumor suppressor genes in human glioma cell lines.
Brain Pathol. 9:469-479(1999)
PubMed=10560660; DOI=10.1097/00005072-199911000-00007
Schmidt E.E., Ichimura K., Goike H.M., Moshref A., Liu L., Collins V.P.
Mutational profile of the PTEN gene in primary human astrocytic tumors and cultivated xenografts.
J. Neuropathol. Exp. Neurol. 58:1170-1183(1999)
PubMed=11414198; DOI=10.1007/s004320000207
Lahm H., Andre S., Hoeflich A., Fischer J.R., Sordat B., Kaltner H., Wolf E., Gabius H.-J.
Comprehensive galectin fingerprinting in a panel of 61 human tumor cell lines by RT-PCR and its implications for diagnostic and therapeutic procedures.
J. Cancer Res. Clin. Oncol. 127:375-386(2001)
PubMed=14614447; DOI=10.1038/sj.onc.1207198
Wischhusen J., Naumann U., Ohgaki H., Rastinejad F., Weller M.
CP-31398, a novel p53-stabilizing agent, induces p53-dependent and p53-independent glioma cell death.
Oncogene 22:8233-8245(2003)
PubMed=14655754; DOI=10.1111/j.1750-3639.2003.tb00479.x
Bahr O., Rieger J., Duffner F., Meyermann R., Weller M., Wick W.
P-glycoprotein and multidrug resistance-associated protein mediate specific patterns of multidrug resistance in malignant glioma cell lines, but not in primary glioma cells.
Brain Pathol. 13:482-494(2003)
PubMed=14961077; DOI=10.1038/sj.onc.1207244
Wang Y.-L., Zhu S.-J., Cloughesy T.F., Liau L.M., Mischel P.S.
p53 disruption profoundly alters the response of human glioblastoma cells to DNA topoisomerase I inhibition.
Oncogene 23:1283-1290(2004)
PubMed=16232199; DOI=10.1111/j.1349-7006.2005.00099.x
Saigusa K., Hashimoto N., Tsuda H., Yokoi S., Maruno M., Yoshimine T., Aoyagi M., Ohno K., Imoto I., Inazawa J.
Overexpressed Skp2 within 5p amplification detected by array-based comparative genomic hybridization is associated with poor prognosis of glioblastomas.
Cancer Sci. 96:676-683(2005)
PubMed=17595512; DOI=10.1159/000104150
Rieger J., Frank B., Weller M., Wick W.
Mechanisms of resistance of human glioma cells to Apo2 ligand/TNF-related apoptosis-inducing ligand.
Cell. Physiol. Biochem. 20:23-34(2007)
PubMed=18445040; DOI=10.1111/j.1742-4658.2008.06452.x
Kikuchi Y., Kakeya T., Nakajima O., Sakai A., Ikeda K., Yamaguchi N., Yamazaki T., Tanamoto K.-i., Matsuda H., Sawada J.-i., Takatori K.
Hypoxia induces expression of a GPI-anchorless splice variant of the prion protein.
FEBS J. 275:2965-2976(2008)
PubMed=19435942; DOI=10.1215/15228517-2009-025
Ichimura K., Pearson D.M., Kocialkowski S., Backlund L.M., Chan R., Jones D.T.W., Collins V.P.
IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas.
Neuro-oncol. 11:341-347(2009)
PubMed=20164919; DOI=10.1038/nature08768
Bignell G.R., Greenman C.D., Davies H., Butler A.P., Edkins S., Andrews J.M., Buck G., Chen L., Beare D., Latimer C., Widaa S., Hinton J., Fahey C., Fu B.-Y., Swamy S., Dalgliesh G.L., Teh B.T., Deloukas P., Yang F.-T., Campbell P.J., Futreal P.A., Stratton M.R.
Signatures of mutation and selection in the cancer genome.
Nature 463:893-898(2010)
PubMed=20215515; DOI=10.1158/0008-5472.CAN-09-3458
Rothenberg S.M., Mohapatra G., Rivera M.N., Winokur D., Greninger P., Nitta M., Sadow P.M., Sooriyakumar G., Brannigan B.W., Ulman M.J., Perera R.M., Wang R., Tam A., Ma X.-J., Erlander M., Sgroi D.C., Rocco J.W., Lingen M.W., Cohen E.E.W., Louis D.N., Settleman J., Haber D.A.
A genome-wide screen for microdeletions reveals disruption of polarity complex genes in diverse human cancers.
Cancer Res. 70:2158-2164(2010)
PubMed=20587068; DOI=10.1186/1471-2407-10-339
Nakabayashi H., Yawata T., Shimizu K.
Anti-invasive and antiangiogenic effects of MMI-166 on malignant glioma cells.
BMC Cancer 10:339.1-339.9(2010)
PubMed=22460905; DOI=10.1038/nature11003
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=22570425; DOI=10.1093/neuonc/nos072
Bady P., Diserens A.-C., Castella V., Kalt S., Heinimann K., Hamou M.-F., Delorenzi M., Hegi M.E.
DNA fingerprinting of glioma cell lines and considerations on similarity measurements.
Neuro-oncol. 14:701-711(2012)
PubMed=25984343; DOI=10.1038/sdata.2014.35
Cowley G.S., Weir B.A., Vazquez F., Tamayo P., Scott J.A., Rusin S., East-Seletsky A., Ali L.D., Gerath W.F.J., Pantel S.E., Lizotte P.H., Jiang G.-Z., Hsiao J., Tsherniak A., Dwinell E., Aoyama S., Okamoto M., Harrington W., Gelfand E.T., Green T.M., Tomko M.J., Gopal S., Wong T.C., Li H.-B., Howell S., Stransky N., Liefeld T., Jang D., Bistline J., Meyers B.H., Armstrong S.A., Anderson K.C., Stegmaier K., Reich M., Pellman D., Boehm J.S., Mesirov J.P., Golub T.R., Root D.E., Hahn W.C.
Parallel genome-scale loss of function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies.
Sci. Data 1:140035-140035(2014)
PubMed=25485619; DOI=10.1038/nbt.3080
Klijn C., Durinck S., Stawiski E.W., Haverty P.M., Jiang Z.-S., Liu H.-B., Degenhardt J., Mayba O., Gnad F., Liu J.-F., Pau G., Reeder J., Cao Y., Mukhyala K., Selvaraj S.K., Yu M.-M., Zynda G.J., Brauer M.J., Wu T.D., Gentleman R.C., Manning G., Yauch R.L., Bourgon R., Stokoe D., Modrusan Z., Neve R.M., de Sauvage F.J., Settleman J., Seshagiri S., Zhang Z.-M.
A comprehensive transcriptional portrait of human cancer cell lines.
Nat. Biotechnol. 33:306-312(2015)
PubMed=25877200; DOI=10.1038/nature14397
Yu M., Selvaraj S.K., Liang-Chu M.M.Y., Aghajani S., Busse M., Yuan J., Lee G., Peale F.V., Klijn C., Bourgon R., Kaminker J.S., Neve R.M.
A resource for cell line authentication, annotation and quality control.
Nature 520:307-311(2015)
PubMed=25894527; DOI=10.1371/journal.pone.0121314
Bausch-Fluck D., Hofmann A., Bock T., Frei A.P., Cerciello F., Jacobs A., Moest H., Omasits U., Gundry R.L., Yoon C., Schiess R., Schmidt A., Mirkowska P., Hartlova A.S., Van Eyk J.E., Bourquin J.-P., Aebersold R., Boheler K.R., Zandstra P.W., Wollscheid B.
A mass spectrometric-derived cell surface protein atlas.
PLoS ONE 10:E0121314-E0121314(2015)
PubMed=26496030; DOI=10.18632/oncotarget.6171
Patil V., Pal J., Somasundaram K.
Elucidating the cancer-specific genetic alteration spectrum of glioblastoma derived cell lines from whole exome and RNA sequencing.
Oncotarget 6:43452-43471(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=27397505; DOI=10.1016/j.cell.2016.06.017
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., Aben N., Goncalves E., Barthorpe S., Lightfoot H., Cokelaer T., Greninger P., van Dyk E., Chang H., de Silva H., Heyn H., Deng X.-M., Egan R.K., Liu Q.-S., Mironenko T., Mitropoulos X., Richardson L., Wang J.-H., Zhang T.-H., Moran S., Sayols S., Soleimani M., Tamborero D., Lopez-Bigas N., Ross-Macdonald P., Esteller M., Gray N.S., Haber D.A., Stratton M.R., Benes C.H., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Garnett M.J.
A landscape of pharmacogenomic interactions in cancer.
Cell 166:740-754(2016)
PubMed=27412690; DOI=10.1074/mcp.M116.060350
Shraibman B., Kadosh D.M., Barnea E., Admon A.
Human leukocyte antigen (HLA) peptides derived from tumor antigens induced by inhibition of DNA methylation for development of drug-facilitated immunotherapy.
Mol. Cell. Proteomics 15:3058-3070(2016)
PubMed=30188626; DOI=10.1134/S1990519X16050072
Kiseleva L.N., Kartashev A.V., Vartanyan N.L., Pinevich A.A., Samoylovich M.P.
Characteristics of A172 and T98G cell lines.
Tsitologiia 58:349-355(2016)
PubMed=28464039; DOI=10.1371/journal.pone.0176683
Hill V.K., Kim J.-S., James C.D., Waldman T.
Correction of PTEN mutations in glioblastoma cell lines via AAV-mediated gene editing.
PLoS ONE 12:E0176683-E0176683(2017)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=30971826; DOI=10.1038/s41586-019-1103-9
Behan F.M., Iorio F., Picco G., Goncalves E., Beaver C.M., Migliardi G., Santos R., Rao Y., Sassi F., Pinnelli M., Ansari R., Harper S., Jackson D.A., McRae R., Pooley R., Wilkinson P., van der Meer D.J., Dow D., Buser-Doepner C.A., Bertotti A., Trusolino L., Stronach E.A., Saez-Rodriguez J., Yusa K., Garnett M.J.
Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens.
Nature 568:511-516(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. III, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=33385022; DOI=10.1016/j.dib.2020.106643
de Sousa J.F., da Silva P., Serafim R.B., Nociti R.P., Moreira C.G., Silva W.A., Valente V.
RNA sequencing data of different grade astrocytoma cell lines.
Data Brief 34:106643.1-106643.13(2021)
PubMed=35839778; DOI=10.1016/j.ccell.2022.06.010
Goncalves E., Poulos R.C., Cai Z.-X., Barthorpe S., Manda S.S., Lucas N., Beck A., Bucio-Noble D., Dausmann M., Hall C., Hecker M., Koh J., Lightfoot H., Mahboob S., Mali I., Morris J., Richardson L., Seneviratne A.J., Shepherd R., Sykes E., Thomas F., Valentini S., Williams S.G., Wu Y.-X., Xavier D., MacKenzie K.L., Hains P.G., Tully B., Robinson P.J., Zhong Q., Garnett M.J., Reddel R.R.
Pan-cancer proteomic map of 949 human cell lines.
Cancer Cell 40:835-849.e8(2022)

 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司