|
产品名称 |
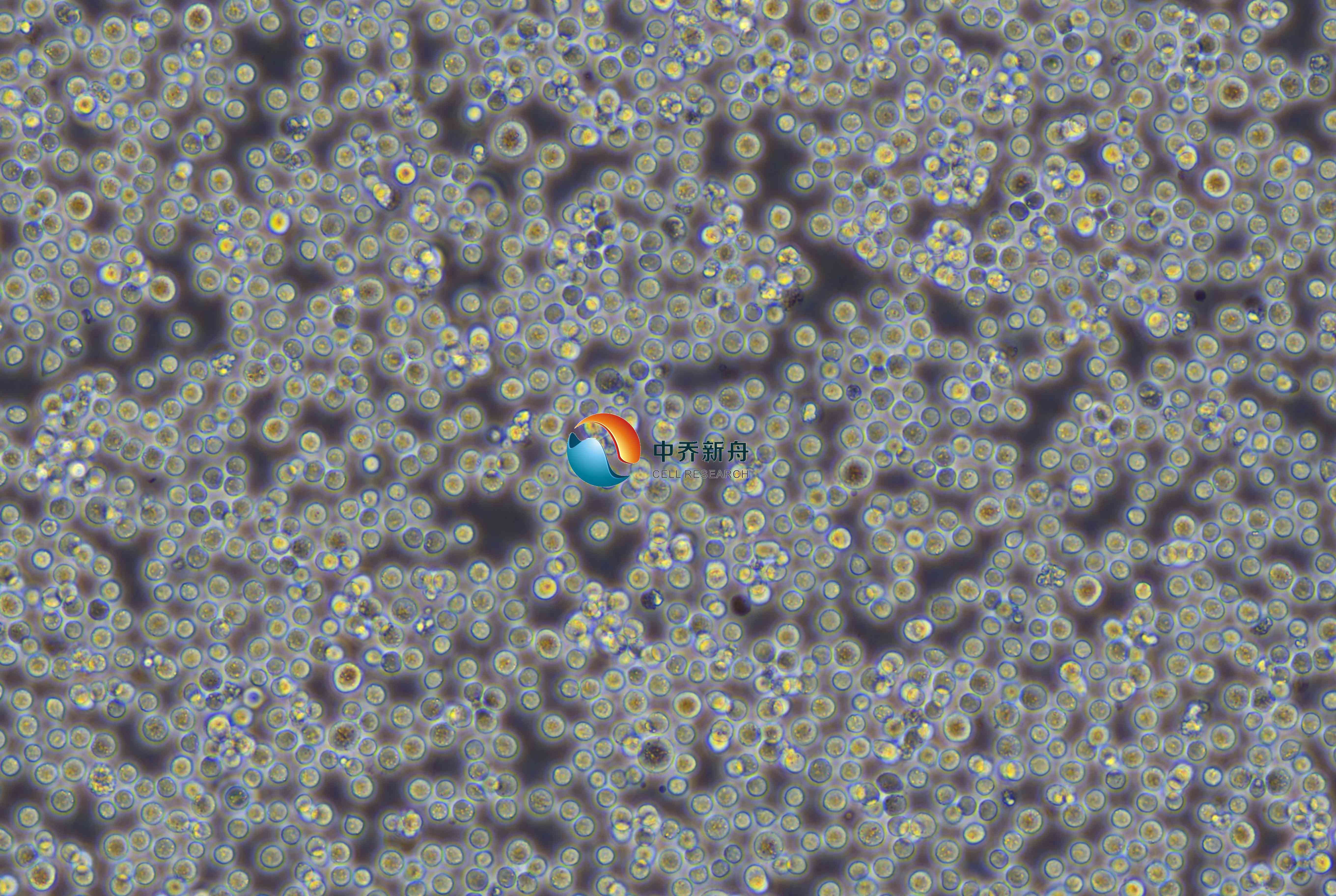
REC1人套细胞淋巴瘤细胞 |
|
货号 |
ZQ1092 |
|
产品介绍 |
REC-1是一种人套细胞淋巴瘤细胞系,源自2003年一位57岁男性的淋巴结中分离出的淋巴母细胞系,该患者患有终末难治性B细胞非霍奇金淋巴瘤。REC-1细胞系是由携带T(11;14)(Q13;Q32)的淋巴瘤建立的,这一染色体易位是套细胞淋巴瘤的特征之一,可用于免疫学研究。 |
|
种属 |
人 |
|
性别/年龄 |
男/57岁 |
|
组织 |
淋巴结 |
|
疾病 |
套细胞淋巴瘤 |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
淋巴母细胞 |
|
生长方式 |
悬浮 |
|
倍增时间 |
大约:38 hours (PubMed=25315077); ~38 hours (ATCC=CRL-3004); ~20-40 hours (DSMZ=ACC-584) |
|
培养基和添加剂 |
RPMI-1640(品牌:中乔新舟 货号:ZQ-200)+10%胎牛血清(中乔新舟 货号:ZQ500-A)+1%P/S(中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
|
|
生物安全等级 |
BSL-1 |
|
培养条件 |
95%空气,5%二氧化碳;37℃ |
|
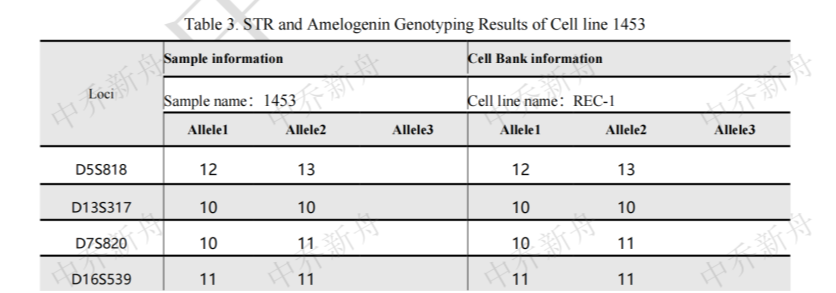
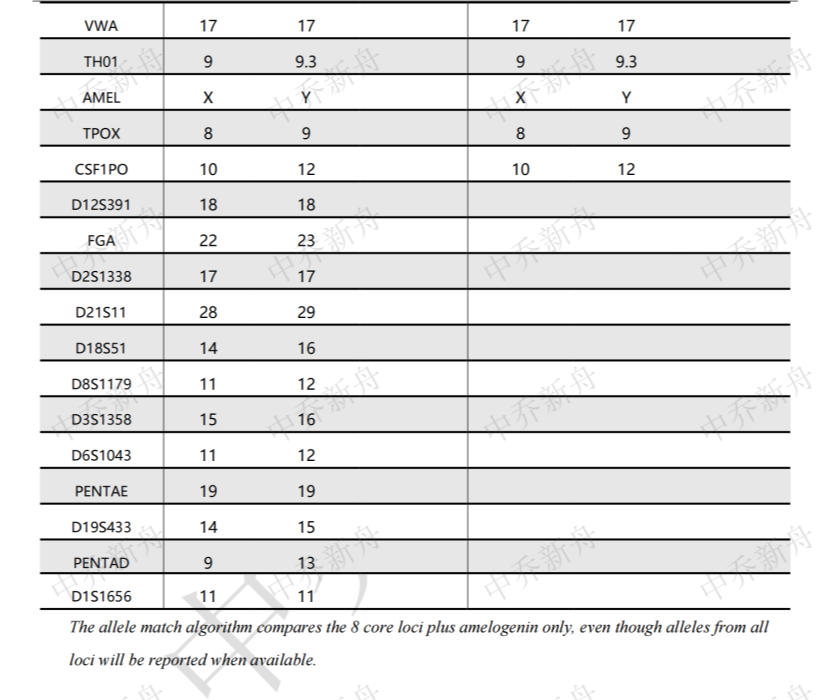
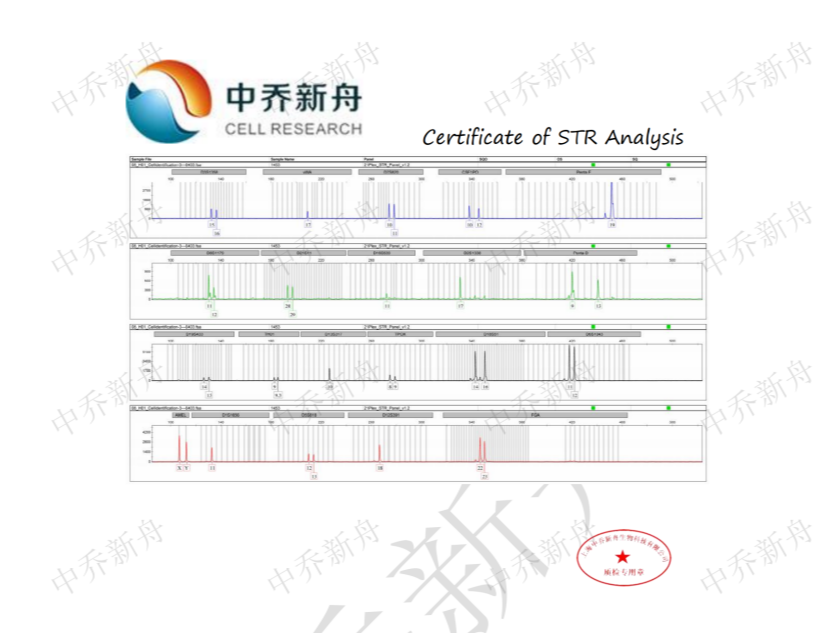
STR位点信息 |
Amelogenin: X,Y CSF1PO :10,12 D2S1338: 17 D3S1358 :15,16 D5S818: 12,13 D7S820: 10,11 D8S1179 :11,12 D13S317: 10 D16S539: 11 D18S51: 14,16 D19S433: 14,15 D21S11 :28,29 FGA :22,23 Penta D: 9,13 Penta E: 7 (PubMed=25877200) :7,19 (DSMZ=ACC-584) :19 (ATCC=CRL-3004) TH01: 9,9.3 TPOX: 8,9 vWA :17 |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ATCC; CRL-3004 ,DSMZ; ACC-584 ,IZSLER; BS TCL 227 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZQ1092 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial )二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |
PubMed=1572647; DOI=10.1016/0888-7543(92)90303-A
Szepetowski P., Simon M.-P., Grosgeorge J., Huebner K., Bastard C., Evans G.A., Tsujimoto Y., Birnbaum D., Theillet C., Gaudray P.
Localization of 11q13 loci with respect to regional chromosomal breakpoints.
Genomics 12:738-744(1992)
PubMed=7504521; DOI=10.1002/gcc.2870080204
Raynaud S.D., Bekri S., Leroux D., Grosgeorge J., Klein B., Bastard C., Gaudray P., Simon M.-P.
Expanded range of 11q13 breakpoints with differing patterns of cyclin D1 expression in B-cell malignancies.
Genes Chromosomes Cancer 8:80-87(1993)
PubMed=8558920
Dirks W.G., Zaborski M., Jager K., Challier C., Shiota M., Quentmeier H., Drexler H.G.
The (2;5)(p23;q35) translocation in cell lines derived from malignant lymphomas: absence of t(2;5) in Hodgkin-analogous cell lines.
Leukemia 10:142-149(1996)
PubMed=14973505; DOI=10.1038/sj.leu.2403315
Prod'homme T., Drenou B., De Ruyffelaere C., Barbieri G., Wiszniewski W., Bastard C., Charron D., Alcaide-Loridan C.
Defective class II transactivator expression in a B lymphoma cell line.
Leukemia 18:832-840(2004)
PubMed=15126345; DOI=10.1158/0008-5472.CAN-03-3773
Ota A., Tagawa H., Karnan S., Tsuzuki S., Karpas A., Kira S., Yoshida Y., Seto M.
Identification and characterization of a novel gene, C13orf25, as a target for 13q31-q32 amplification in malignant lymphoma.
Cancer Res. 64:3087-3095(2004)
PubMed=16448697; DOI=10.1016/j.leukres.2005.11.013
Camps J., Salaverria I., Garcia M.J., Prat E., Bea S., Pole J.C.M., Hernandez L., Del Rey J., Cigudosa J.C., Bernues M., Caldas C., Colomer D., Miro R., Campo E.
Genomic imbalances and patterns of karyotypic variability in mantle-cell lymphoma cell lines.
Leuk. Res. 30:923-934(2006)
PubMed=16960149; DOI=10.1182/blood-2006-06-026500
Mestre-Escorihuela C., Rubio-Moscardo F., Richter J.A., Siebert R., Climent J., Fresquet V., Beltran E., Agirre X., Marugan I., Marin M., Rosenwald A., Sugimoto K.-j., Wheat L.M., Karran E.L., Garcia J.F., Sanchez-Verde L., Prosper F., Staudt L.M., Pinkel D., Dyer M.J.S., Martinez-Climent J.A.
Homozygous deletions localize novel tumor suppressor genes in B-cell lymphomas.
Blood 109:271-280(2007)
PubMed=17332242; DOI=10.1182/blood-2006-11-057208
Pinyol M., Bea S., Pla L., Ribrag V., Bosq J., Rosenwald A., Campo E., Jares P.
Inactivation of RB1 in mantle-cell lymphoma detected by nonsense-mediated mRNA decay pathway inhibition and microarray analysis.
Blood 109:5422-5429(2007)
PubMed=20215515; DOI=10.1158/0008-5472.CAN-09-3458
Rothenberg S.M., Mohapatra G., Rivera M.N., Winokur D., Greninger P., Nitta M., Sadow P.M., Sooriyakumar G., Brannigan B.W., Ulman M.J., Perera R.M., Wang R., Tam A., Ma X.-J., Erlander M., Sgroi D.C., Rocco J.W., Lingen M.W., Cohen E.E.W., Louis D.N., Settleman J., Haber D.A.
A genome-wide screen for microdeletions reveals disruption of polarity complex genes in diverse human cancers.
Cancer Res. 70:2158-2164(2010)
PubMed=21746927; DOI=10.1073/pnas.1018941108
Beltran E., Fresquet V., Martinez-Useros J., Richter-Larrea J.A., Sagardoy A., Sesma I., Almada L.L., Montes-Moreno S., Siebert R., Gesk S., Calasanz M.J., Malumbres R., Rieger M., Prosper F., Lossos I.S., Piris M.A., Fernandez-Zapico M.E., Martinez-Climent J.A.
A cyclin-D1 interaction with BAX underlies its oncogenic role and potential as a therapeutic target in mantle cell lymphoma.
Proc. Natl. Acad. Sci. U.S.A. 108:12461-12466(2011)
PubMed=22460905; DOI=10.1038/nature11003
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=24362935; DOI=10.1038/nm.3435
Rahal R., Frick M., Romero R., Korn J.M., Kridel R., Chan F.C., Meissner B., Bhang H.-E.C., Ruddy D., Kauffmann A., Farsidjani A., Derti A., Rakiec D., Naylor T., Pfister E., Kovats S., Kim S., Dietze K., Dorken B., Steidl C., Tzankov A., Hummel M., Monahan J., Morrissey M.P., Fritsch C., Sellers W.R., Cooke V.G., Gascoyne R.D., Lenz G., Stegmeier F.
Pharmacological and genomic profiling identifies NF-kappaB-targeted treatment strategies for mantle cell lymphoma.
Nat. Med. 20:87-92(2014)
PubMed=25960936; DOI=10.4161/21624011.2014.954893
Boegel S., Lower M., Bukur T., Sahin U., Castle J.C.
A catalog of HLA type, HLA expression, and neo-epitope candidates in human cancer cell lines.
OncoImmunology 3:e954893.1-e954893.12(2014)
PubMed=25315077; DOI=10.3109/10428194.2014.970548
Fogli L.K., Williams M.E., Connors J.M., Reid Y.A., Brown K., O'Connor O.A.
Development and characterization of a Mantle Cell Lymphoma Cell Bank in the American Type Culture Collection.
Leuk. Lymphoma 56:2114-2122(2015)
PubMed=25355872; DOI=10.1128/JVI.02570-14
Cao S.-B., Strong M.J., Wang X., Moss W.N., Concha M., Lin Z., O'Grady T., Baddoo M., Fewell C., Renne R., Flemington E.K.
High-throughput RNA sequencing-based virome analysis of 50 lymphoma cell lines from the Cancer Cell Line Encyclopedia project.
J. Virol. 89:713-729(2015)
PubMed=25485619; DOI=10.1038/nbt.3080
Klijn C., Durinck S., Stawiski E.W., Haverty P.M., Jiang Z.-S., Liu H.-B., Degenhardt J., Mayba O., Gnad F., Liu J.-F., Pau G., Reeder J., Cao Y., Mukhyala K., Selvaraj S.K., Yu M.-M., Zynda G.J., Brauer M.J., Wu T.D., Gentleman R.C., Manning G., Yauch R.L., Bourgon R., Stokoe D., Modrusan Z., Neve R.M., de Sauvage F.J., Settleman J., Seshagiri S., Zhang Z.-M.
A comprehensive transcriptional portrait of human cancer cell lines.
Nat. Biotechnol. 33:306-312(2015)
PubMed=25688540; DOI=10.1002/cyto.a.22643
Maiga S., Brosseau C., Descamps G., Dousset C., Gomez-Bougie P., Chiron D., Menoret E., Kervoelen C., Vie H., Cesbron A., Moreau-Aubry A., Amiot M., Pellat-Deceunynck C.
A simple flow cytometry-based barcode for routine authentication of multiple myeloma and mantle cell lymphoma cell lines.
Cytometry A 87:285-288(2015)
PubMed=25877200; DOI=10.1038/nature14397
Yu M., Selvaraj S.K., Liang-Chu M.M.Y., Aghajani S., Busse M., Yuan J., Lee G., Peale F.V., Klijn C., Bourgon R., Kaminker J.S., Neve R.M.
A resource for cell line authentication, annotation and quality control.
Nature 520:307-311(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=28196595; DOI=10.1016/j.ccell.2017.01.005
Li J., Zhao W., Akbani R., Liu W.-B., Ju Z.-L., Ling S.-Y., Vellano C.P., Roebuck P., Yu Q.-H., Eterovic A.K., Byers L.A., Davies M.A., Deng W.-L., Gopal Y.N.V., Chen G., von Euw E.M., Slamon D.J., Conklin D., Heymach J.V., Gazdar A.F., Minna J.D., Myers J.N., Lu Y.-L., Mills G.B., Liang H.
Characterization of human cancer cell lines by reverse-phase protein arrays.
Cancer Cell 31:225-239(2017)
PubMed=29666304; DOI=10.1158/1078-0432.CCR-17-3004
Pham L.V., Huang S.-J., Zhang H., Zhang J., Bell T., Zhou S.-H., Pogue E., Ding Z.-Y., Lam L., Westin J., Davis R.E., Young K.H., Medeiros L.J., Ford R.J. Jr., Nomie K., Zhang L., Wang M.
Strategic therapeutic targeting to overcome venetoclax resistance in aggressive B-cell lymphomas.
Clin. Cancer Res. 24:3967-3980(2018)
PubMed=30285677; DOI=10.1186/s12885-018-4840-5
Tan K.-T., Ding L.-W., Sun Q.-Y., Lao Z.-T., Chien W., Ren X., Xiao J.-F., Loh X.-Y., Xu L., Lill M., Mayakonda A., Lin D.-C., Yang H., Koeffler H.P.
Profiling the B/T cell receptor repertoire of lymphocyte derived cell lines.
BMC Cancer 18:940.1-940.13(2018)
PubMed=30629668; DOI=10.1371/journal.pone.0210404
Uphoff C.C., Pommerenke C., Denkmann S.A., Drexler H.G.
Screening human cell lines for viral infections applying RNA-Seq data analysis.
PLoS ONE 14:E0210404-E0210404(2019)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. III, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=31160637; DOI=10.1038/s41598-019-44491-x
Quentmeier H., Pommerenke C., Dirks W.G., Eberth S., Koeppel M., MacLeod R.A.F., Nagel S., Steube K., Uphoff C.C., Drexler H.G.
The LL-100 panel: 100 cell lines for blood cancer studies.
Sci. Rep. 9:8218-8218(2019)
PubMed=31978347; DOI=10.1016/j.cell.2019.12.023
Nusinow D.P., Szpyt J., Ghandi M., Rose C.M., McDonald E.R. III, Kalocsay M., Jane-Valbuena J., Gelfand E.T., Schweppe D.K., Jedrychowski M.P., Golji J., Porter D.A., Rejtar T., Wang Y.K., Kryukov G.V., Stegmeier F., Erickson B.K., Garraway L.A., Sellers W.R., Gygi S.P.
Quantitative proteomics of the Cancer Cell Line Encyclopedia.
Cell 180:387-402.e16(2020)
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司