|
产品名称
|
KYSE-510人食管鳞状细胞癌细胞
|
|
货号
|
ZQ1124
|
|
产品介绍
|
KYSE-510是一种人食管鳞状细胞癌细胞系,最初是从一名 67 岁女性接受顺铂和放疗后颈段食管切除的外膜中度浸润的高分化食管鳞状细胞癌建立。描述为携带TP53突变和扩增的MYC、HST1、细胞周期蛋白D1,并在裸鼠中诱导分化良好的肿瘤。
|
|
种属
|
人
|
|
性别/年龄
|
女/67岁
|
|
组织
|
食管
|
|
疾病
|
人食管鳞状细胞癌
|
|
细胞类型
|
肿瘤细胞
|
|
形态学
|
上皮
|
|
生长方式
|
贴壁
|
|
培养基和添加剂
|
RPMI-1640(中乔新舟 货号:ZQ-200)+F-12 基础培养基(中乔新舟 货号:ZQ-400)(基础1:1混合)+10%胎牛血清(中乔新舟 货号:ZQ500-A)+1%P/S(中乔新舟 货号:CSP006)
|
|
推荐完全培养基货号
|
ZM1124
|
|
生物安全等级
|
BSL-1
|
|
培养条件
|
95%空气,5%二氧化碳;37℃
|
|
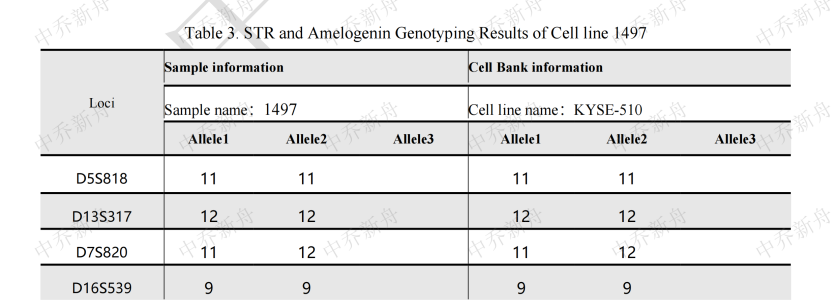
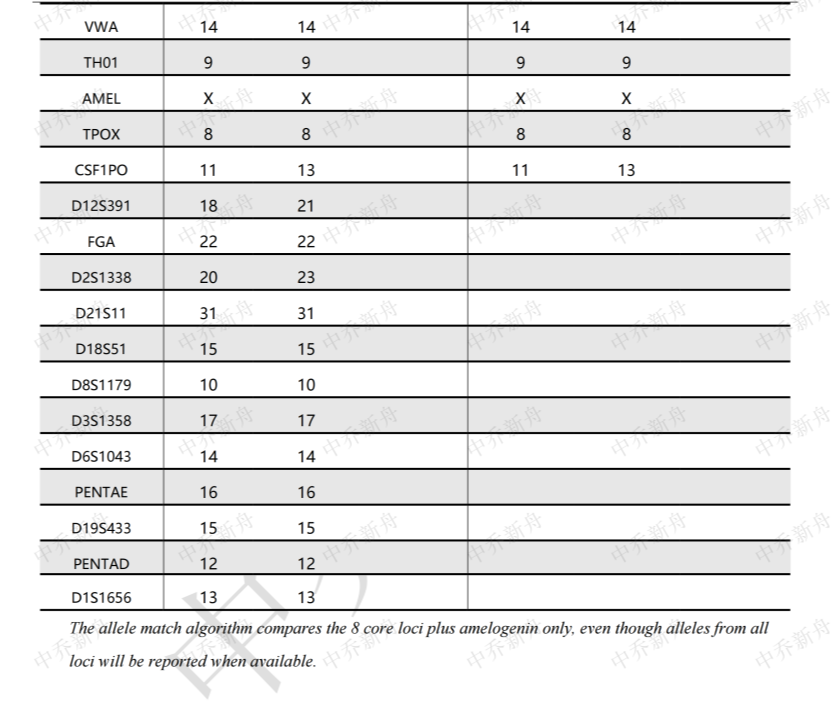
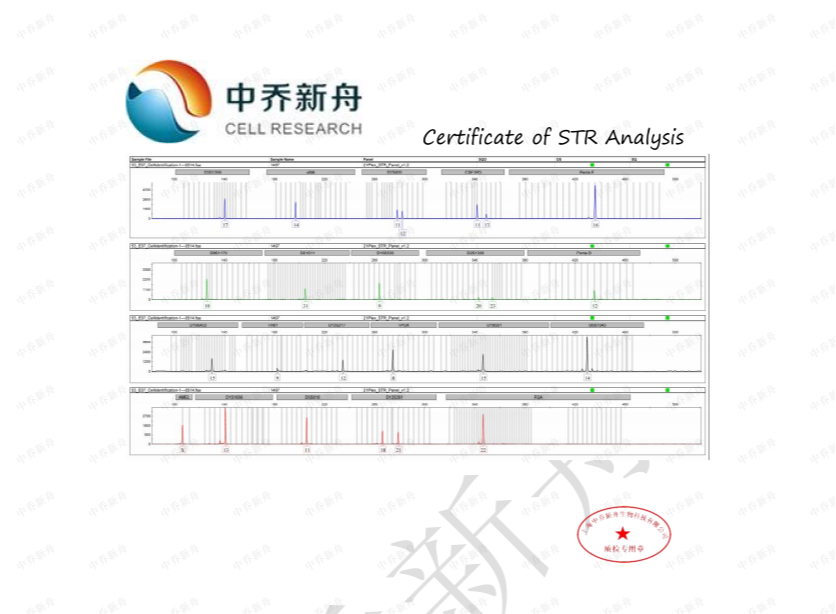
STR位点信息
|
Amelogenin :X
CSF1PO: 11 (DSMZ=ACC-374)
:11,13 (Cosmic-CLP=907321; JCRB=JCRB1436; PubMed=25877200)
D2S1338: 20,23
D3S1358 :17
D5S818 :11
D7S820 :11,12
D8S1179 :10
D13S317 :12
D16S539 :9
D18S51: 15
D19S433: 15
D21S11 :31
FGA: 22
Penta D :12
Penta E :16
TH01 :9
TPOX :8
vWA :14
|
|
抗原表达/受体表达
|
***
|
|
基因表达
|
***
|
|
保藏机构
|
DSMZ; ACC-374 JCRB; JCRB1436
|
|
供应限制
|
仅供科研使用
|
|
货号
|
ZQ1124
|
|
发货规格
|
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial )二选一
|
|
发货形式
|
活细胞:常温运输;冻存管:干冰运输
|
|
储存温度
|
活细胞:培养箱;冻存管:液氮罐
|
|
产地
|
中国
|
|
供应限制
|
仅供科研使用
|
PubMed=1728357; DOI=10.1002/1097-0142(19920115)69:2<277::AID-CNCR2820690202>3.0.CO;2-C
Shimada Y., Imamura M., Wagata T., Yamaguchi N., Tobe T.
Characterization of 21 newly established esophageal cancer cell lines.
Cancer 69:277-284(1992)
PubMed=7913084; DOI=10.1002/ijc.2910580224
Kanda Y., Nishiyama Y., Shimada Y., Imamura M., Nomura H., Hiai H., Fukumoto M.
Analysis of gene amplification and overexpression in human esophageal-carcinoma cell lines.
Int. J. Cancer 58:291-297(1994)
PubMed=8575860; DOI=10.1002/(SICI)1097-0215(19960126)65:3<372::AID-IJC16>3.0.CO;2-C
Tanaka H., Shibagaki I., Shimada Y., Wagata T., Imamura M., Ishizaki K.
Characterization of p53 gene mutations in esophageal squamous cell carcinoma cell lines: increased frequency and different spectrum of mutations from primary tumors.
Int. J. Cancer 65:372-376(1996)
PubMed=9033652; DOI=10.1002/(SICI)1097-0215(19970207)70:4<437::AID-IJC11>3.0.CO;2-C
Tanaka H., Shimada Y., Imamura M., Shibagaki I., Ishizaki K.
Multiple types of aberrations in the p16 (INK4a) and the p15(INK4b) genes in 30 esophageal squamous-cell-carcinoma cell lines.
Int. J. Cancer 70:437-442(1997)
PubMed=11092977; DOI=10.1111/j.1349-7006.2000.tb00895.x
Pimkhaokham A., Shimada Y., Fukuda Y., Kurihara N., Imoto I., Yang Z.-Q., Imamura M., Nakamura Y., Amagasa T., Inazawa J.
Nonrandom chromosomal imbalances in esophageal squamous cell carcinoma cell lines: possible involvement of the ATF3 and CENPF genes in the 1q32 amplicon.
Jpn. J. Cancer Res. 91:1126-1133(2000)
PubMed=15172977; DOI=10.1158/0008-5472.CAN-04-0172
Sonoda I., Imoto I., Inoue J., Shibata T., Shimada Y., Chin K., Imamura M., Amagasa T., Gray J.W., Hirohashi S., Inazawa J.
Frequent silencing of low density lipoprotein receptor-related protein 1B (LRP1B) expression by genetic and epigenetic mechanisms in esophageal squamous cell carcinoma.
Cancer Res. 64:3741-3747(2004)
PubMed=16045545; DOI=10.1111/j.0959-9673.2005.00431.x
Ban S., Michikawa Y., Ishikawa K.-i., Sagara M., Watanabe K., Shimada Y., Inazawa J., Imai T.
Radiation sensitivities of 31 human oesophageal squamous cell carcinoma cell lines.
Int. J. Exp. Pathol. 86:231-240(2005)
PubMed=20164919; DOI=10.1038/nature08768
Bignell G.R., Greenman C.D., Davies H., Butler A.P., Edkins S., Andrews J.M., Buck G., Chen L., Beare D., Latimer C., Widaa S., Hinton J., Fahey C., Fu B.-Y., Swamy S., Dalgliesh G.L., Teh B.T., Deloukas P., Yang F.-T., Campbell P.J., Futreal P.A., Stratton M.R.
Signatures of mutation and selection in the cancer genome.
Nature 463:893-898(2010)
PubMed=20215515; DOI=10.1158/0008-5472.CAN-09-3458
Rothenberg S.M., Mohapatra G., Rivera M.N., Winokur D., Greninger P., Nitta M., Sadow P.M., Sooriyakumar G., Brannigan B.W., Ulman M.J., Perera R.M., Wang R., Tam A., Ma X.-J., Erlander M., Sgroi D.C., Rocco J.W., Lingen M.W., Cohen E.E.W., Louis D.N., Settleman J., Haber D.A.
A genome-wide screen for microdeletions reveals disruption of polarity complex genes in diverse human cancers.
Cancer Res. 70:2158-2164(2010)
PubMed=22460905; DOI=10.1038/nature11003
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=25984343; DOI=10.1038/sdata.2014.35
Cowley G.S., Weir B.A., Vazquez F., Tamayo P., Scott J.A., Rusin S., East-Seletsky A., Ali L.D., Gerath W.F.J., Pantel S.E., Lizotte P.H., Jiang G.-Z., Hsiao J., Tsherniak A., Dwinell E., Aoyama S., Okamoto M., Harrington W., Gelfand E.T., Green T.M., Tomko M.J., Gopal S., Wong T.C., Li H.-B., Howell S., Stransky N., Liefeld T., Jang D., Bistline J., Meyers B.H., Armstrong S.A., Anderson K.C., Stegmaier K., Reich M., Pellman D., Boehm J.S., Mesirov J.P., Golub T.R., Root D.E., Hahn W.C.
Parallel genome-scale loss of function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies.
Sci. Data 1:140035-140035(2014)
PubMed=25485619; DOI=10.1038/nbt.3080
Klijn C., Durinck S., Stawiski E.W., Haverty P.M., Jiang Z.-S., Liu H.-B., Degenhardt J., Mayba O., Gnad F., Liu J.-F., Pau G., Reeder J., Cao Y., Mukhyala K., Selvaraj S.K., Yu M.-M., Zynda G.J., Brauer M.J., Wu T.D., Gentleman R.C., Manning G., Yauch R.L., Bourgon R., Stokoe D., Modrusan Z., Neve R.M., de Sauvage F.J., Settleman J., Seshagiri S., Zhang Z.-M.
A comprehensive transcriptional portrait of human cancer cell lines.
Nat. Biotechnol. 33:306-312(2015)
PubMed=25877200; DOI=10.1038/nature14397
Yu M., Selvaraj S.K., Liang-Chu M.M.Y., Aghajani S., Busse M., Yuan J., Lee G., Peale F.V., Klijn C., Bourgon R., Kaminker J.S., Neve R.M.
A resource for cell line authentication, annotation and quality control.
Nature 520:307-311(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=27397505; DOI=10.1016/j.cell.2016.06.017
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., Aben N., Goncalves E., Barthorpe S., Lightfoot H., Cokelaer T., Greninger P., van Dyk E., Chang H., de Silva H., Heyn H., Deng X.-M., Egan R.K., Liu Q.-S., Mironenko T., Mitropoulos X., Richardson L., Wang J.-H., Zhang T.-H., Moran S., Sayols S., Soleimani M., Tamborero D., Lopez-Bigas N., Ross-Macdonald P., Esteller M., Gray N.S., Haber D.A., Stratton M.R., Benes C.H., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Garnett M.J.
A landscape of pharmacogenomic interactions in cancer.
Cell 166:740-754(2016)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=30971826; DOI=10.1038/s41586-019-1103-9
Behan F.M., Iorio F., Picco G., Goncalves E., Beaver C.M., Migliardi G., Santos R., Rao Y., Sassi F., Pinnelli M., Ansari R., Harper S., Jackson D.A., McRae R., Pooley R., Wilkinson P., van der Meer D.J., Dow D., Buser-Doepner C.A., Bertotti A., Trusolino L., Stronach E.A., Saez-Rodriguez J., Yusa K., Garnett M.J.
Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens.
Nature 568:511-516(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. III, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=31978347; DOI=10.1016/j.cell.2019.12.023
Nusinow D.P., Szpyt J., Ghandi M., Rose C.M., McDonald E.R. III, Kalocsay M., Jane-Valbuena J., Gelfand E.T., Schweppe D.K., Jedrychowski M.P., Golji J., Porter D.A., Rejtar T., Wang Y.K., Kryukov G.V., Stegmeier F., Erickson B.K., Garraway L.A., Sellers W.R., Gygi S.P.
Quantitative proteomics of the Cancer Cell Line Encyclopedia.
Cell 180:387-402.e16(2020)
PubMed=35839778; DOI=10.1016/j.ccell.2022.06.010
Goncalves E., Poulos R.C., Cai Z.-X., Barthorpe S., Manda S.S., Lucas N., Beck A., Bucio-Noble D., Dausmann M., Hall C., Hecker M., Koh J., Lightfoot H., Mahboob S., Mali I., Morris J., Richardson L., Seneviratne A.J., Shepherd R., Sykes E., Thomas F., Valentini S., Williams S.G., Wu Y.-X., Xavier D., MacKenzie K.L., Hains P.G., Tully B., Robinson P.J., Zhong Q., Garnett M.J., Reddel R.R.
Pan-cancer proteomic map of 949 human cell lines.
Cancer Cell 40:835-849.e8(2022)
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司