|
产品名称 |
KATO III人胃癌细胞 |
|
货号 |
ZQ0238 |
|
产品介绍 |
从一名55岁男性患者的胸腔积液中体外建立了人胃癌细胞系KATO-III。
KATO-III 细胞系是一种源自低分化腺癌转移部位的人类胃癌模型。这些细胞广泛用于胃癌研究,特别是用于研究驱动肿瘤进展、耐药性和转移的分子机制。KATO-III 细胞表现出非整倍体核型,其特征是多种染色体异常,这导致了它们的侵袭性癌症表型。它们明显缺乏 p53,这一特征通常与致瘤性增加和化疗反应改变有关,使其成为研究 p53 在胃癌中作用的宝贵工具。 注意事项:
KATO III细胞为不均匀大小圆细胞,细胞形态为悬浮细胞和贴壁细胞共存生长。大部分圆细胞可以吹打,部分贴壁细胞需消化1min。 |
|
种属 |
人 |
|
性别/年龄 |
男/55岁 |
|
组织 |
胃;来源于转移部位:胸腔积液、锁骨上及腋窝淋巴结和道格拉斯氏陷凹 |
|
疾病 |
胃癌 |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
球形 |
|
生长方式 |
混合;贴壁和悬浮 |
|
倍增时间 |
大约32~41小时 |
|
培养基和添加剂 |
IMDM(中乔新舟 货号:ZQ-900)+20%胎牛血清(中乔新舟 货号:ZQ500-A)+1%双抗(中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
|
|
生物安全等级 |
BSL-1 |
|
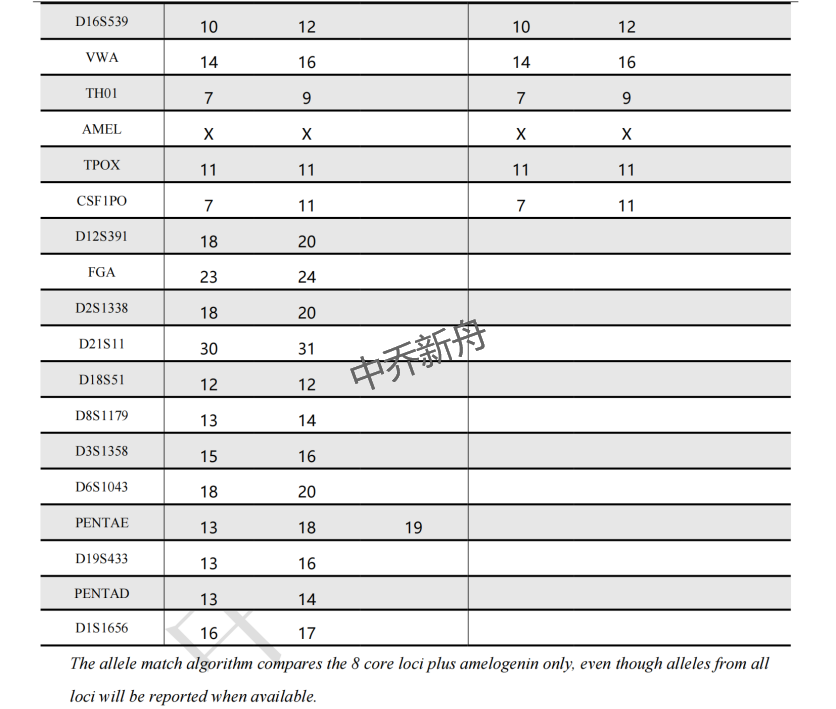
STR位点信息 |
Amelogenin: X
D3S1358: 15,16
|
|
培养条件 |
95%空气,5%二氧化碳;37℃ |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ATCC; HTB-103 ECACC; 86093004 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZM0238 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial)二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |


| 货号 | 产品名称 | 规格 | 价格 | 指令 |
| ZQ0055 | BGC-823人胃癌细胞(hela污染) | 1x10^6 Cells/Vial | ¥1400.00 | 放入购物车 》 |
| ZQ0059 | MGC-803人胃癌细胞(hela污染) | 1x10^6 Cells/Vial | ¥1400.00 | 放入购物车 》 |
| ZQ0060 | NCI-N87人胃癌细胞(STR鉴定)[细胞+500ml专培套餐促销] | 1x10^6 Cells/Vial | ¥980 | 放入购物车 》 |
| ZQ0062 | SGC-7901人胃癌细胞(hela污染) | 1x10^6 Cells/Vial | ¥1700 | 放入购物车 》 |
| ZQ0112 | BGC-803人胃癌细胞(暂不提供) | 1x10^6 Cells/Vial | ¥1400.00 | 放入购物车 》 |
| ZQ0187 | MFC小鼠胃癌细胞(种属鉴定)[细胞+500ml专培套餐促销] | 1x10^6 Cells/Vial | ¥980.00 | 放入购物车 》 |
| ZQ0192 | HGC-27人胃癌细胞(未分化)(STR鉴定)[细胞+500ml专培套餐促销] | 1x10^6 Cells/Vial | ¥980.00 | 放入购物车 》 |
| ZQ0239 | SNU-1人胃癌细胞(STR鉴定) | 1x10^6 Cells/Vial | ¥1600.00 | 放入购物车 》 |
| ZQ0457 | MKN-45人胃癌细胞 (STR鉴定)[细胞+500ml专培套餐促销] | 1x10^6 Cells/Vial | ¥980.00 | 放入购物车 》 |
| ZQ506 | 支原体清除试剂盒 | 1ml | ¥600.00 | 放入购物车 》 |
| 1 | 慢病毒介导基因沉默或过表达 | ¥询价 | 放入购物车 》 | |
| ZQ500-A | 优级胎牛血清 | 500ml | ¥2580(开学促销) | 放入购物车 》 |
| ZQ-900 | IMDM 基础培养基 | 500ml | ¥96.00 | 放入购物车 》 |
| ZQ-902 | IMDM 完全培养基(20%FBS) | 500ml | ¥650.00 | 放入购物车 》 |
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司