|
产品名称 |
AsPC-1人转移胰腺癌细胞 |
|
货号 |
ZQ0254 |
|
产品介绍 |
一个胰腺癌病人的腹水中的细胞移植到裸鼠后建立了这个细胞株。 注意事项: 该细胞对血清质量敏感,建议用户使用优质血清进行培养。如遇细胞死亡较多或生长缓慢,可选用我司提供的该细胞完全培养液。
AsPC-1细胞生长呈不规则形态。细胞消化传代后细胞完全贴壁生长需要24-48小时,48小时才能换液操作。 |
|
种属 |
人 |
|
性别/年龄 |
女/62岁 |
|
组织 |
胰腺;来源于转移部位:腹水 |
|
疾病 |
腺癌 |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
上皮细胞 |
|
生长方式 |
贴壁 |
|
倍增时间 |
大约38~46小时 |
|
培养基和添加剂 |
RPMI-1640年(中乔新舟 货号:ZQ-200)+10%胎牛血清(中乔新舟货号:ZQ0500)+1%双抗(中乔新舟货号:CSP006) |
|
推荐完全培养基货号 |
|
|
生物安全等级 |
BSL-1 |
|
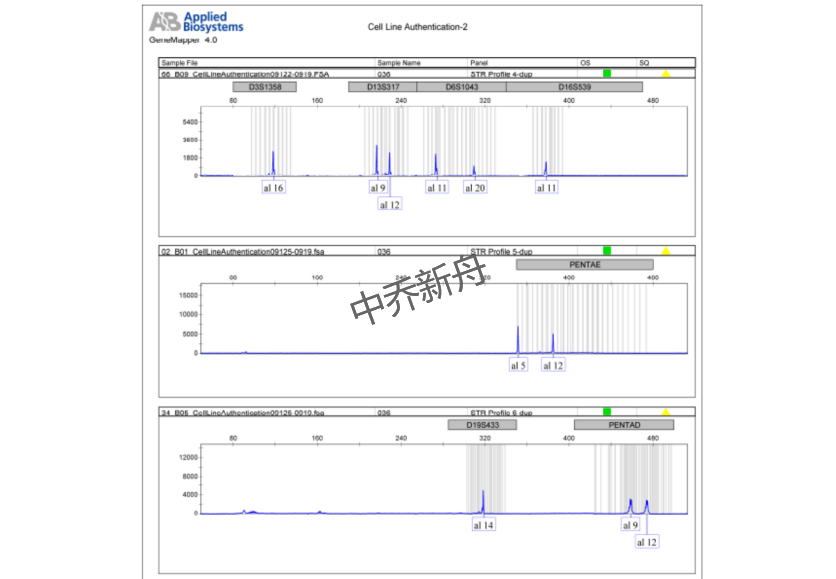
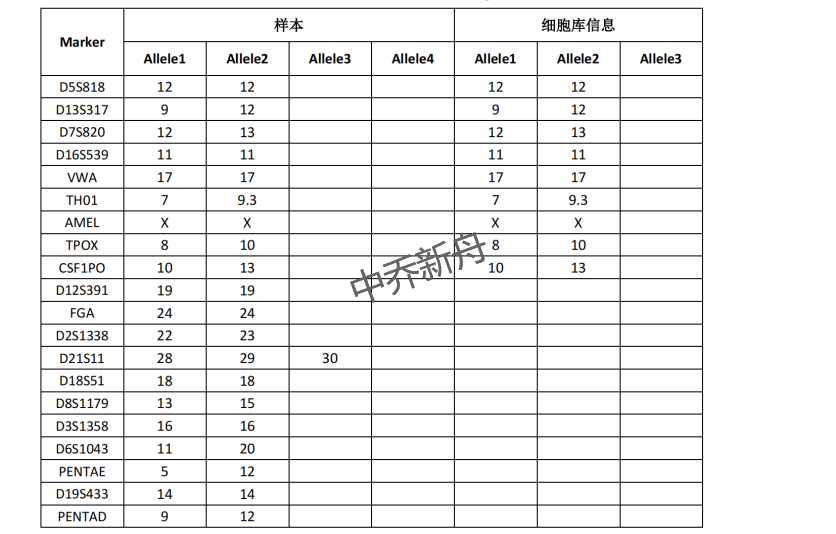
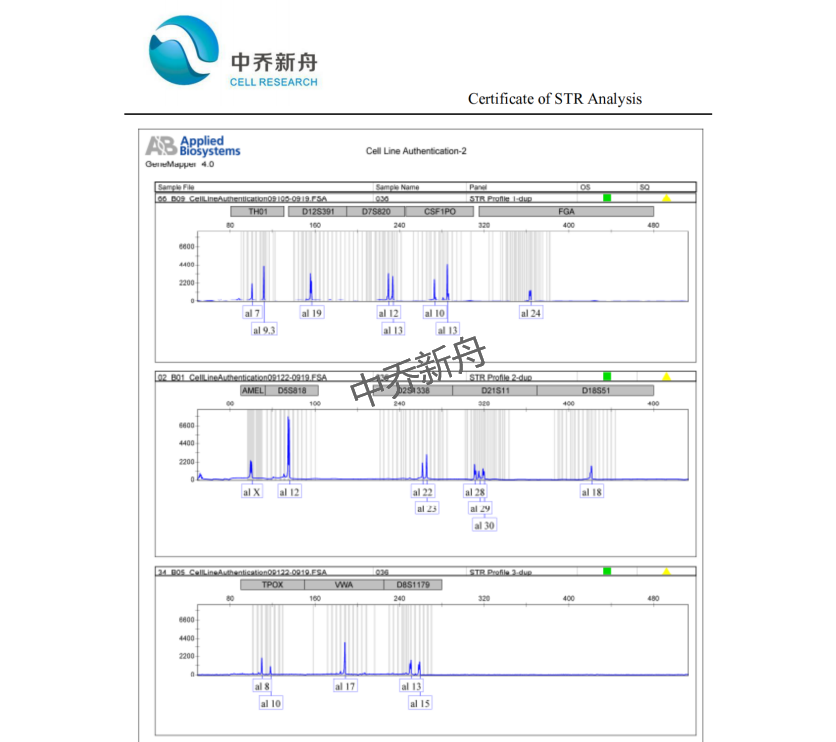
力量值位点信息 |
釉原蛋白:X CSF1PO: 10,13 D13S317: 9,12 D16S539: 11 D5S818: 12 D7S820: 12,13 TH01: 7,9.3 TPOX: 8,10 vWA: 17 |
|
培养条件 |
95%空气,5%二氧化碳;37℃ |
|
抗原表达/受体表达 |
|
|
基因表达 |
|
|
保藏机构 |
ATCC;BCRC CRL-1682;60494 ECACC;96020930 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZM0254 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1毫升冻存管*2支(细胞量约为1x10^6 Cells/Vial)二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |

| 货号 | 产品名称 | 规格 | 价格 | 指令 |
| ZQ0065 | PANC-1人胰腺癌细胞(STR鉴定)[细胞+500ml专培套餐促销] | 1x10^6 Cells/Vial | ¥980 | 放入购物车 》 |
| ZQ0252 | CFPAC-1人胰腺癌细胞(STR鉴定)[细胞+500ml专培套餐促销] | 1x10^6 Cells/Vial | ¥980.00 | 放入购物车 》 |
| ZQ0253 | SW 1990人胰腺癌细胞(STR鉴定) | 1x10^6 Cells/Vial | ¥4000.00 | 放入购物车 》 |
| ZQ0454 | MIA PaCa-2人胰腺癌细胞(STR鉴定)[细胞+500ml专培套餐促销] | 1x10^6 Cells/Vial | ¥980.00 | 放入购物车 》 |
| ZQ505 | 支原体检测试剂盒(买二送一) | 1ml/100T | ¥800.00 | 放入购物车 》 |
| ZQ506 | 支原体清除试剂盒 | 1ml | ¥600.00 | 放入购物车 》 |
| A1001-1 | 国产转染试剂 | 1ml | ¥800.00 | 放入购物车 》 |
| 1 | 慢病毒介导基因沉默或过表达 | ¥询价 | 放入购物车 》 | |
| 6 | 绿/红色荧光蛋白标记 | ¥询价 | 放入购物车 》 | |
| ZQ-200 | RPMI-1640基础培养基 | 500ml | ¥56.00 | 放入购物车 》 |
| ZQ-201 | RPMI-1640完全培养基(10%FBS) | 500ml | ¥350.00 | 放入购物车 》 |
| ZQ500-A | 优级胎牛血清 | 500ml | ¥2580(开学促销) | 放入购物车 》 |
| JSFW-134 | 人源细胞STR鉴定服务 | ¥900.00 | 放入购物车 》 |
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司