|
产品名称 |
MDA-MB-435S人黑素瘤细胞 |
|
货号 |
ZQ0374 |
|
产品介绍 |
该细胞系最初被描述为 R. Cailleau 等人于 1976 年分离的亲本 MDA-MB-435 菌株的纺锤形变体。来自一名患有转移性乳腺导管腺癌的 31 岁女性的胸腔积液。 然而,最近的研究对亲本细胞系 MDA-MB-435 以及延伸的 HTB-129 的起源产生了疑问。对细胞的基因表达分析产生了微阵列,其中 MDA-MB-435 与黑色素瘤来源的细胞系聚集在一起。 此后的更多研究证实了 MDA-MB-435 的黑素细胞起源,ATCC 对此做出了回应,对该细胞系的身份进行了自己的调查。据报道,MDA-MB-435 交叉污染的细胞系是 M14 黑色素瘤细胞系 [PubMed ID:12354931 和 PubMed ID:17004106]。 |
|
种属 |
人 |
|
性别/年龄 |
女/31岁 |
|
组织 |
先前描述为:乳腺/乳房;来源于转移部位:胸腔积液 |
|
疾病 |
无色素性黑色素瘤 |
|
细胞类型 |
黑色素细胞,黑色素瘤 |
|
形态学 |
纺锤形 |
|
生长方式 |
贴壁 |
|
倍增时间 |
大约48~72小时 |
|
培养基和添加剂(默认) |
DMEM (中乔新舟 货号:ZQ-100)+10%FBS(中乔新舟 货号:ZQ0500)+1%双抗(中乔新舟 货号:CSP006) 注意默认方案培养条件:95%空气,5%CO2;37℃ |
|
培养方案B(可选) |
L-15(中乔新舟 货号:ZQ-1100)+10%FBS(中乔新舟 货号:ZQ500-A)+1%P/S(中乔新舟 货号:CSP006)+0.01mg/mL牛胰岛素+0.01mg/ml谷胱甘肽 注意B方案培养条件:100%空气;37℃ |
|
推荐默认完全培养基货号 |
|
|
B方案完全培养基货号 |
|
|
生物安全等级 |
BSL-1 |
|
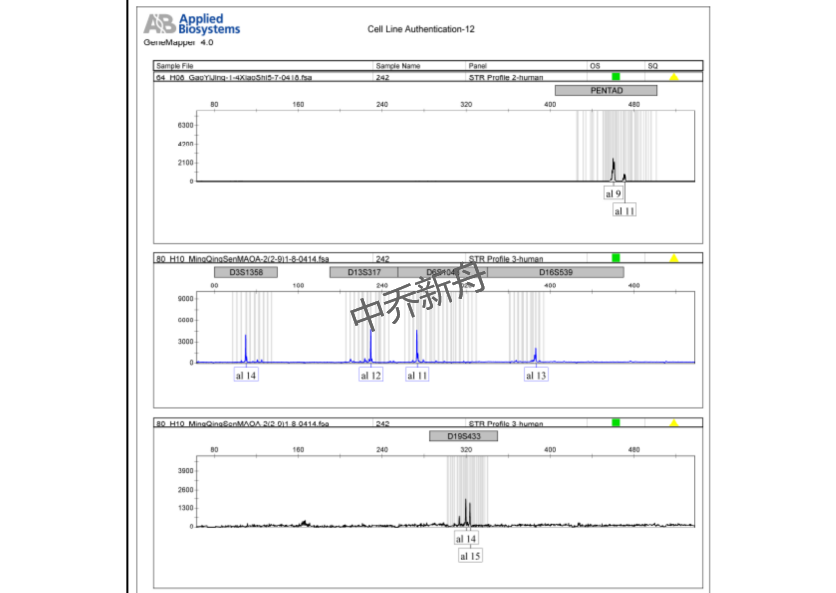
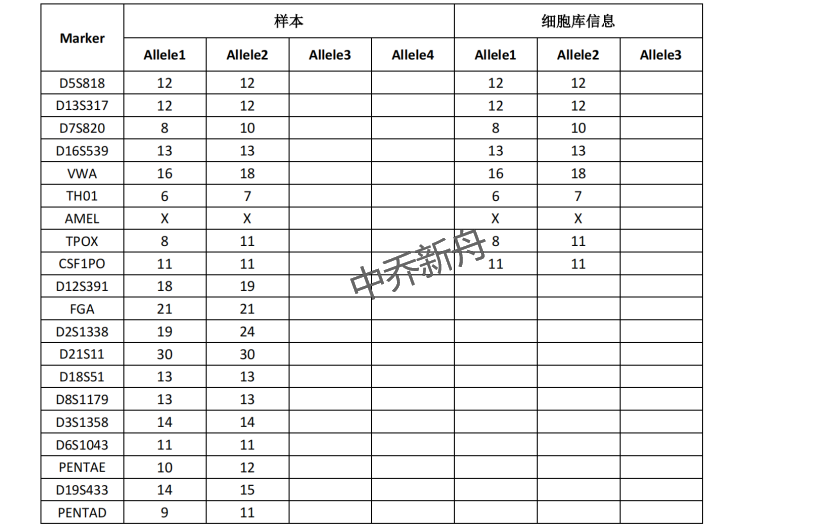
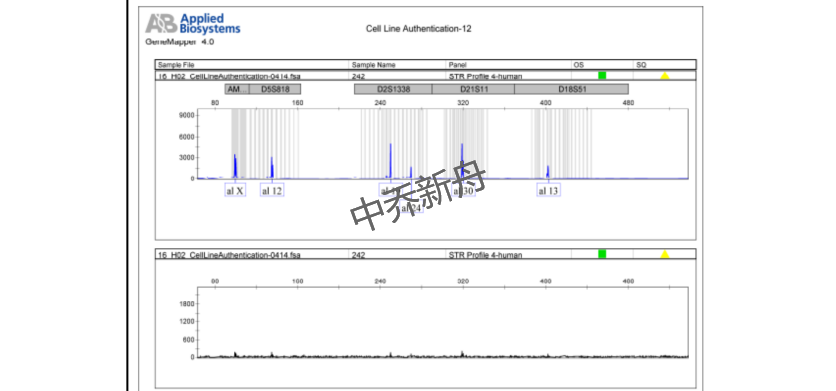
STR位点信息 |
Amelogenin: X CSF1PO: 11 D13S317: 12 D16S539: 13 D5S818: 12 D7S820: 8,10 TH01: 6,7 TPOX: 8,11 vWA: 16,18 |
|
默认方案培养条件 |
95%空气,5%CO2;37℃ |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ATCC; HTB-129 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZQ0374 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial)二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |


| 货号 | 产品名称 | 规格 | 价格 | 指令 |
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司