|
产品名称 |
U251人神经胶质瘤细胞 |
|
货号 |
ZQ0053 |
|
产品介绍 |
U251是一种人神经胶质瘤细胞系,源自一位75岁男性患者的胶质母细胞瘤组织。胶质母细胞瘤是一种常见的、恶性程度很高的原发性脑肿瘤,通常发生在星形胶质细胞中。U251在神经胶质瘤的研究中非常重要,它们具有快速的体外增殖能力、高度的侵袭性,并且携带多种与肿瘤发生和发展相关的基因突变,如TP53和EGFR突变。是神经胶质瘤研究的重要工具,它尽管存在局限性,但通过结合多种研究方法和技术,科学家们可以进一步揭示神经胶质瘤的生物学机制,并开发出更有效的治疗策略。 |
|
种属 |
人 |
|
性别/年龄 |
男/75岁 |
|
组织 |
脑组织 |
|
疾病 |
大脑顶叶;疾病:星形胶质细胞瘤(III-IV级) |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
成纤维细胞样,多形型, |
|
生长方式 |
贴壁 |
|
倍增时间 |
~24小时 |
|
培养基和添加剂 |
DMEM高糖(中乔新舟 货号:ZQ-100)+10%胎牛血清(中乔新舟 货号:ZQ0500)+1%P/S(中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
|
|
生物安全等级 |
BSL-1 |
|
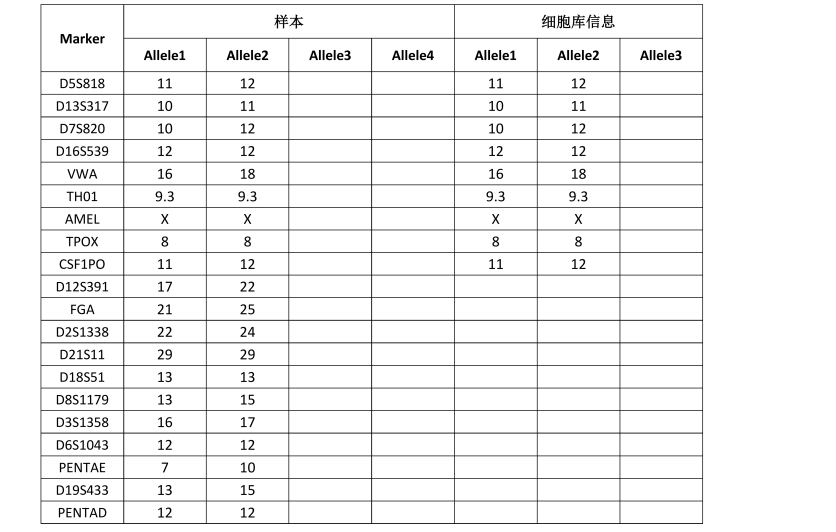
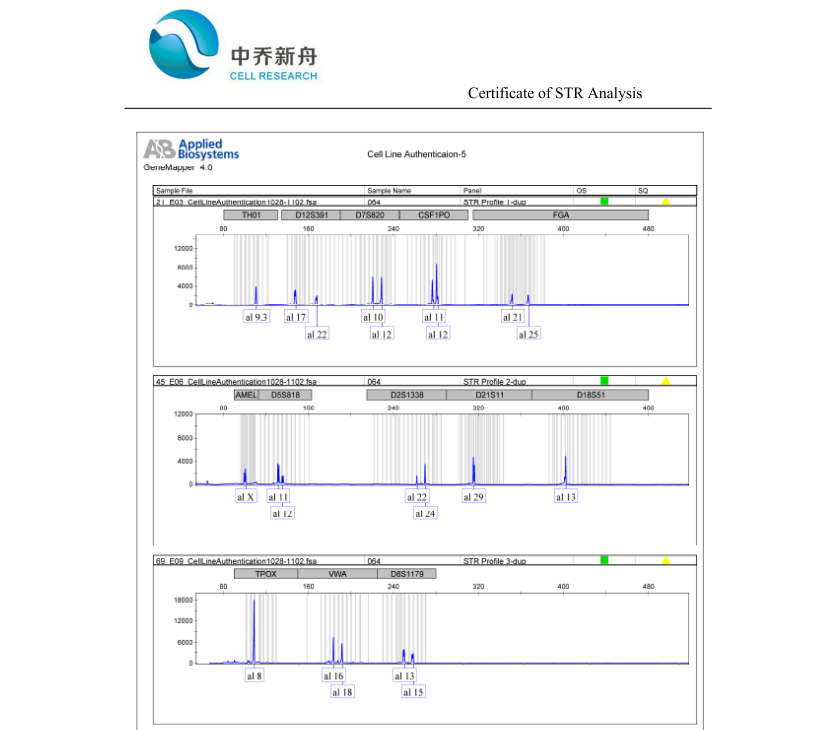
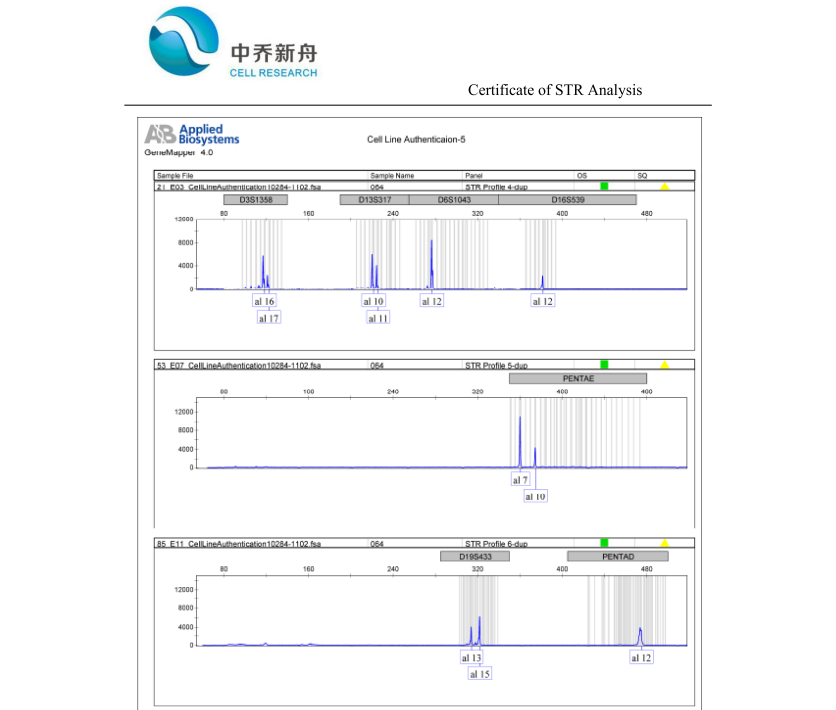
STR位点信息 |
Amelogenin: X; |
|
培养条件 |
95%空气,5%二氧化碳;37℃ |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
ECACC; 09063001 |
|
供应限制 |
仅供科研使用 |
|
货号 |
ZQ0053 |
|
发货规格 |
活细胞:T25培养瓶*1瓶或者1ml 冻存管*2支(细胞量约为1x10^6 Cells/Vial)二选一 |
|
发货形式 |
活细胞:常温运输;冻存管:干冰运输 |
|
储存温度 |
活细胞:培养箱;冻存管:液氮罐 |
|
产地 |
中国 |
|
供应限制 |
仅供科研使用 |
原文链接: https://www.mdpi.com/1089476
论文标题: Cyclic arginine-glycine-aspartic acid-modified red blood cells for drug delivery: synthesis and in vitro evaluation
DOI: 10.1016/j.jpha.2021.06.003
发表时间: 2021-06-12
期刊: Journal of Pharmaceutical Analysis
影响因子: 4.769
货号: ZQ0053
产品名称: U251 glioma cells
原文链接: https://www.sciencedirect.com/science/article/pii/S2095177921000630
PubMed=4134536; DOI=10.1002/ijc.2910120215
Westermark B.
The deficient density-dependent growth control of human malignant glioma cells and virus-transformed glia-like cells in culture.
Int. J. Cancer 12:438-451(1973)
PubMed=4369403; DOI=10.1002/1097-0142(197408)34:2<274::AID-CNCR2820340209>3.0.CO;2-4
Wahlstrom T., Linder E., Saksela E., Westermark B.
Tumor-specific membrane antigens in established cell lines from gliomas.
Cancer 34:274-279(1974)
DOI=10.1007/978-1-4757-1647-4_7
Ponten J.
Neoplastic human glia cells in culture.
(In book chapter) Human tumor cells in vitro; Fogh J. (eds.); pp.175-206; Springer; New York; USA (1975)
PubMed=608846; DOI=10.1111/j.1601-5223.1978.tb01267.x
Mark J., Westermark B., Ponten J., Hugosson R.
Banding patterns in human glioma cell lines.
Hereditas 87:243-260(1977)
PubMed=833871; DOI=10.1093/jnci/58.2.209
Fogh J., Wright W.C., Loveless J.D.
Absence of HeLa cell contamination in 169 cell lines derived from human tumors.
J. Natl. Cancer Inst. 58:209-214(1977)
PubMed=77569; DOI=10.1111/j.1399-0039.1978.tb01259.x
Espmark J.A., Ahlqvist-Roth L., Sarne L., Persson A.
Tissue typing of cells in culture. III. HLA antigens of established human cell lines. Attempts at typing by the mixed hemadsorption technique.
Tissue Antigens 11:279-286(1978)
PubMed=450131; DOI=10.1038/279797a0
Day R.S. 3rd, Ziolkowski C.H.J.
Human brain tumour cell strains with deficient host-cell reactivation of N-methyl-N'-nitro-N-nitrosoguanidine-damaged adenovirus 5.
Nature 279:797-799(1979)
PubMed=6260907; DOI=10.1097/00005072-198105000-00001
Bigner D.D., Bigner S.H., Ponten J., Westermark B., Mahaley M.S. Jr., Ruoslahti E., Herschman H., Eng L.F., Wikstrand C.J.
Heterogeneity of genotypic and phenotypic characteristics of fifteen permanent cell lines derived from human gliomas.
J. Neuropathol. Exp. Neurol. 40:201-229(1981)
PubMed=6582512; DOI=10.1073/pnas.81.2.568; PMCID=PMC344720
Mattes M.J., Cordon-Cardo C., Lewis J.L. Jr., Old L.J., Lloyd K.O.
Cell surface antigens of human ovarian and endometrial carcinoma defined by mouse monoclonal antibodies.
Proc. Natl. Acad. Sci. U.S.A. 81:568-572(1984)
PubMed=3518877; DOI=10.3109/07357908609038260
Fogh J.
Human tumor lines for cancer research.
Cancer Invest. 4:157-184(1986)
PubMed=3675803
Bigner S.H., Bjerkvig R., Laerum O.D., Muhlbaier L.H., Bigner D.D.
DNA content and chromosomes in permanent cultured cell lines derived from malignant human gliomas.
Anal. Quant. Cytol. Histol. 9:435-444(1987)
PubMed=3335022
Alley M.C., Scudiero D.A., Monks A., Hursey M.L., Czerwinski M.J., Fine D.L., Abbott B.J., Mayo J.G., Shoemaker R.H., Boyd M.R.
Feasibility of drug screening with panels of human tumor cell lines using a microculture tetrazolium assay.
Cancer Res. 48:589-601(1988)
PubMed=2041050; DOI=10.1093/jnci/83.11.757
Monks A., Scudiero D.A., Skehan P., Shoemaker R.H., Paull K.D., Vistica D.T., Hose C.D., Langley J., Cronise P., Vaigro-Wolff A., Gray-Goodrich M., Campbell H., Mayo J.G., Boyd M.R.
Feasibility of a high-flux anticancer drug screen using a diverse panel of cultured human tumor cell lines.
J. Natl. Cancer Inst. 83:757-766(1991)
PubMed=7693337
Li H., Hamou M.-F., de Tribolet N., Jaufeerally R., Hofmann M., Diserens A.-C., Van Meir E.G.
Variant CD44 adhesion molecules are expressed in human brain metastases but not in glioblastomas.
Cancer Res. 53:5345-5349(1993)
PubMed=7763724
Satoh M., Takeuchi M.
Cross-contamination of cell lines as revealed by DNA fingerprinting in the IFO animal cell bank.
Res. Commun. Inst. Ferment. 16:18-23(1993)
PubMed=8104691
Pershouse M.A., Stubblefield E., Hadi A., Killary A.M., Yung W.-K.A., Steck P.A.
Analysis of the functional role of chromosome 10 loss in human glioblastomas.
Cancer Res. 53:5043-5050(1993)
DOI=10.1016/B978-0-12-333530-2.50005-8
Nister M., Westermark B.
Human glioma cell lines.
(In book chapter) Atlas of human tumor cell lines; Hay R.J., Park J.-G., Gazdar A.F. (eds.); pp.17-42; Academic Press; New York; USA (1994)
PubMed=8878451; DOI=10.1111/j.1349-7006.1996.tb02118.x; PMCID=PMC5921198
Zhang S., Endo S., Koga H., Ichikawa T., Feng X., Onda K., Washiyama K., Kumanishi T.
A comparative study of glioma cell lines for p16, p15, p53 and p21 gene alterations.
Jpn. J. Cancer Res. 87:900-907(1996)
PubMed=9090379; DOI=10.1038/ng0497-356
Steck P.A., Pershouse M.A., Jasser S.A., Yung W.-K.A., Lin H., Ligon A.H., Langford L.A., Baumgard M.L., Hattier T., Davis T., Frye C., Hu R., Swedlund B., Teng D.H.-F., Tavtigian S.V.
Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers.
Nat. Genet. 15:356-362(1997)
PubMed=9220028; DOI=10.1016/S0303-7207(97)00080-4
Sharif T.R., Luo W., Sharif M.
Functional expression of bombesin receptor in most adult and pediatric human glioblastoma cell lines; role in mitogenesis and in stimulating the mitogen-activated protein kinase pathway.
Mol. Cell. Endocrinol. 130:119-130(1997)
PubMed=9230885; DOI=10.1093/jnci/89.14.1036
Gomez-Manzano C., Fueyo J., Kyritsis A.P., McDonnell T.J., Steck P.A., Levin V.A., Yung W.-K.A.
Characterization of p53 and p21 functional interactions in glioma cells en route to apoptosis.
J. Natl. Cancer Inst. 89:1036-1044(1997)
PubMed=9614553; DOI=10.1038/sj.gt.3300605
Tada M., Sakuma S., Iggo R.D., Saya H., Sawamura Y., Fujiwara T., Roth J.A.
Monitoring adenoviral p53 transduction efficiency by yeast functional assay.
Gene Ther. 5:339-344(1998)
PubMed=9842975; DOI=10.1002/(SICI)1097-0215(19981218)79:6<640::AID-IJC15>3.0.CO;2-z
Weller M., Rieger J., Grimmel C., Van Meir E.G., De Tribolet N., Krajewski S., Reed J.C., von Deimling A., Dichgans J.
Predicting chemoresistance in human malignant glioma cells: the role of molecular genetic analyses.
Int. J. Cancer 79:640-644(1998)
DOI=10.1007/0-306-46861-1_11
Ali-Osman F.
Brain tumors.
(In book chapter) Human cell culture. Vol. 2. Cancer cell lines part 2; Masters J.R.W., Palsson B.O. (eds.); pp.167-184; Kluwer Academic Publishers; New York; USA (1999)
PubMed=10416987; DOI=10.1111/j.1750-3639.1999.tb00536.x; PMCID=PMC8098486
Ishii N., Maier D., Merlo A., Tada M., Sawamura Y., Diserens A.-C., Van Meir E.G.
Frequent co-alterations of TP53, p16/CDKN2A, p14ARF, PTEN tumor suppressor genes in human glioma cell lines.
Brain Pathol. 9:469-479(1999)
PubMed=10551321; DOI=10.1111/j.1349-7006.1999.tb00838.x; PMCID=PMC5926156
Zhang S.-J., Endo S., Ichikawa T., Yoshimura J., Onda K., Tanaka R., Washiyama K., Kumanishi T.
Rare-type mutations of MMAC1 tumor suppressor gene in human glioma cell lines and their tumors of origin.
Jpn. J. Cancer Res. 90:934-941(1999)
PubMed=10560660; DOI=10.1097/00005072-199911000-00007
Schmidt E.E., Ichimura K., Goike H.M., Moshref A., Liu L., Collins V.P.
Mutational profile of the PTEN gene in primary human astrocytic tumors and cultivated xenografts.
J. Neuropathol. Exp. Neurol. 58:1170-1183(1999)
PubMed=10700174; DOI=10.1038/73432
Ross D.T., Scherf U., Eisen M.B., Perou C.M., Rees C., Spellman P.T., Iyer V.R., Jeffrey S.S., van de Rijn M., Waltham M.C., Pergamenschikov A., Lee J.C.F., Lashkari D., Shalon D., Myers T.G., Weinstein J.N., Botstein D., Brown P.O.
Systematic variation in gene expression patterns in human cancer cell lines.
Nat. Genet. 24:227-235(2000)
PubMed=14614447; DOI=10.1038/sj.onc.1207198
Wischhusen J., Naumann U., Ohgaki H., Rastinejad F., Weller M.
CP-31398, a novel p53-stabilizing agent, induces p53-dependent and p53-independent glioma cell death.
Oncogene 22:8233-8245(2003)
PubMed=14655754; DOI=10.1111/j.1750-3639.2003.tb00479.x; PMCID=PMC8095903
Bahr O., Rieger J., Duffner F., Meyermann R., Weller M., Wick W.
P-glycoprotein and multidrug resistance-associated protein mediate specific patterns of multidrug resistance in malignant glioma cell lines, but not in primary glioma cells.
Brain Pathol. 13:482-494(2003)
PubMed=15748285; DOI=10.1186/1479-5876-3-11; PMCID=PMC555742
Adams S., Robbins F.-M., Chen D., Wagage D., Holbeck S.L., Morse H.C. 3rd, Stroncek D., Marincola F.M.
HLA class I and II genotype of the NCI-60 cell lines.
J. Transl. Med. 3:11.1-11.8(2005)
PubMed=15900046; DOI=10.1093/jnci/dji133
Mashima T., Oh-hara T., Sato S., Mochizuki M., Sugimoto Y., Yamazaki K., Hamada J.-i., Tada M., Moriuchi T., Ishikawa Y., Kato Y., Tomoda H., Yamori T., Tsuruo T.
p53-defective tumors with a functional apoptosome-mediated pathway: a new therapeutic target.
J. Natl. Cancer Inst. 97:765-777(2005)
PubMed=16232199; DOI=10.1111/j.1349-7006.2005.00099.x; PMCID=PMC11159392
Saigusa K., Hashimoto N., Tsuda H., Yokoi S., Maruno M., Yoshimine T., Aoyagi M., Ohno K., Imoto I., Inazawa J.
Overexpressed Skp2 within 5p amplification detected by array-based comparative genomic hybridization is associated with poor prognosis of glioblastomas.
Cancer Sci. 96:676-683(2005)
PubMed=16391870; DOI=10.3892/or.15.2.463
Castigli E., Sciaccaluga M., Schiavoni G., Brozzi F., Fabiani R., Gorello P., Gianfranceschi G.L.
GL15 and U251 glioblastoma-derived human cell lines are peculiarly susceptible to induction of mitotic death by very low concentrations of okadaic acid.
Oncol. Rep. 15:463-470(2006)
PubMed=16697959; DOI=10.1016/j.ccr.2006.03.030
Lee J., Kotliarova S., Kotliarov Y., Li A.-G., Su Q., Donin N.M., Pastorino S., Purow B.W., Christopher N., Zhang W., Park J.K., Fine H.A.
Tumor stem cells derived from glioblastomas cultured in bFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines.
Cancer Cell 9:391-403(2006)
PubMed=17088437; DOI=10.1158/1535-7163.MCT-06-0433; PMCID=PMC2705832
Ikediobi O.N., Davies H.R., Bignell G.R., Edkins S., Stevens C., O'Meara S., Santarius T., Avis T., Barthorpe S., Brackenbury L., Buck G., Butler A.P., Clements J., Cole J., Dicks E., Forbes S., Gray K., Halliday K., Harrison R., Hills K., Hinton J., Hunter C., Jenkinson A., Jones D., Kosmidou V., Lugg R., Menzies A., Mironenko T., Parker A., Perry J., Raine K.M., Richardson D., Shepherd R., Small A., Smith R., Solomon H., Stephens P.J., Teague J.W., Tofts C., Varian J., Webb T., West S., Widaa S., Yates A., Reinhold W.C., Weinstein J.N., Stratton M.R., Futreal P.A., Wooster R.
Mutation analysis of 24 known cancer genes in the NCI-60 cell line set.
Mol. Cancer Ther. 5:2606-2612(2006)
PubMed=17595512; DOI=10.1159/000104150
Rieger J., Frank B., Weller M., Wick W.
Mechanisms of resistance of human glioma cells to Apo2 ligand/TNF-related apoptosis-inducing ligand.
Cell. Physiol. Biochem. 20:23-34(2007)
PubMed=19372543; DOI=10.1158/1535-7163.MCT-08-0921; PMCID=PMC4020356
Lorenzi P.L., Reinhold W.C., Varma S., Hutchinson A.A., Pommier Y., Chanock S.J., Weinstein J.N.
DNA fingerprinting of the NCI-60 cell line panel.
Mol. Cancer Ther. 8:713-724(2009)
PubMed=19435942; DOI=10.1215/15228517-2009-025; PMCID=PMC2743214
Ichimura K., Pearson D.M., Kocialkowski S., Backlund L.M., Chan R., Jones D.T.W., Collins V.P.
IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas.
Neuro-oncol. 11:341-347(2009)
PubMed=20215515; DOI=10.1158/0008-5472.CAN-09-3458; PMCID=PMC2881662
Rothenberg S.M., Mohapatra G., Rivera M.N., Winokur D., Greninger P., Nitta M., Sadow P.M., Sooriyakumar G., Brannigan B.W., Ulman M.J., Perera R.M., Wang R., Tam A., Ma X.-J., Erlander M., Sgroi D.C., Rocco J.W., Lingen M.W., Cohen E.E.W., Louis D.N., Settleman J., Haber D.A.
A genome-wide screen for microdeletions reveals disruption of polarity complex genes in diverse human cancers.
Cancer Res. 70:2158-2164(2010)
PubMed=20593219; DOI=10.1007/s11060-010-0283-9
Blough M.D., Beauchamp D.C., Westgate M.R., Kelly J.J.P., Cairncross J.G.
Effect of aberrant p53 function on temozolomide sensitivity of glioma cell lines and brain tumor initiating cells from glioblastoma.
J. Neurooncol. 102:1-7(2011)
PubMed=22068913; DOI=10.1073/pnas.1111840108; PMCID=PMC3219108
Gillet J.-P., Calcagno A.M., Varma S., Marino M., Green L.J., Vora M.I., Patel C., Orina J.N., Eliseeva T.A., Singal V., Padmanabhan R., Davidson B., Ganapathi R., Sood A.K., Rueda B.R., Ambudkar S.V., Gottesman M.M.
Redefining the relevance of established cancer cell lines to the study of mechanisms of clinical anti-cancer drug resistance.
Proc. Natl. Acad. Sci. U.S.A. 108:18708-18713(2011)
PubMed=22336246; DOI=10.1016/j.bmc.2012.01.017
Kong D.-X., Yamori T.
JFCR39, a panel of 39 human cancer cell lines, and its application in the discovery and development of anticancer drugs.
Bioorg. Med. Chem. 20:1947-1951(2012)
PubMed=22347499; DOI=10.1371/journal.pone.0031628; PMCID=PMC3276511
Ruan X.-Y., Kocher J.-P.A., Pommier Y., Liu H.-F., Reinhold W.C.
Mass homozygotes accumulation in the NCI-60 cancer cell lines as compared to HapMap trios, and relation to fragile site location.
PLoS ONE 7:E31628-E31628(2012)
PubMed=22384151; DOI=10.1371/journal.pone.0032096; PMCID=PMC3285665
Lee J.-S., Kim Y.K., Kim H.J., Hajar S., Tan Y.L., Kang N.-Y., Ng S.H., Yoon C.N., Chang Y.-T.
Identification of cancer cell-line origins using fluorescence image-based phenomic screening.
PLoS ONE 7:E32096-E32096(2012)
PubMed=22460905; DOI=10.1038/nature11003; PMCID=PMC3320027
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=22570425; DOI=10.1093/neuonc/nos072; PMCID=PMC3367844
Bady P., Diserens A.-C., Castella V., Kalt S., Heinimann K., Hamou M.-F., Delorenzi M., Hegi M.E.
DNA fingerprinting of glioma cell lines and considerations on similarity measurements.
Neuro-oncol. 14:701-711(2012)
PubMed=22628656; DOI=10.1126/science.1218595; PMCID=PMC3526189
Jain M., Nilsson R., Sharma S., Madhusudhan N., Kitami T., Souza A.L., Kafri R., Kirschner M.W., Clish C.B., Mootha V.K.
Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation.
Science 336:1040-1044(2012)
PubMed=23856246; DOI=10.1158/0008-5472.CAN-12-3342; PMCID=PMC4893961
Abaan O.D., Polley E.C., Davis S.R., Zhu Y.-L.J., Bilke S., Walker R.L., Pineda M.A., Gindin Y., Jiang Y., Reinhold W.C., Holbeck S.L., Simon R.M., Doroshow J.H., Pommier Y., Meltzer P.S.
The exomes of the NCI-60 panel: a genomic resource for cancer biology and systems pharmacology.
Cancer Res. 73:4372-4382(2013)
PubMed=23933261; DOI=10.1016/j.celrep.2013.07.018
Moghaddas Gholami A., Hahne H., Wu Z.-X., Auer F.J., Meng C., Wilhelm M., Kuster B.
Global proteome analysis of the NCI-60 cell line panel.
Cell Rep. 4:609-620(2013)
PubMed=24279929; DOI=10.1186/2049-3002-1-20; PMCID=PMC4178206
Dolfi S.C., Chan L.L.-Y., Qiu J., Tedeschi P.M., Bertino J.R., Hirshfield K.M., Oltvai Z.N., Vazquez A.
The metabolic demands of cancer cells are coupled to their size and protein synthesis rates.
Cancer Metab. 1:20.1-20.13(2013)
PubMed=24670534; DOI=10.1371/journal.pone.0092047; PMCID=PMC3966786
Varma S., Pommier Y., Sunshine M., Weinstein J.N., Reinhold W.C.
High resolution copy number variation data in the NCI-60 cancer cell lines from whole genome microarrays accessible through CellMiner.
PLoS ONE 9:E92047-E92047(2014)
PubMed=24810477; DOI=10.1002/cam4.219; PMCID=PMC4303149
Torsvik A., Stieber D., Enger P.O., Golebiewska A., Molven A., Svendsen A., Westermark B., Niclou S.P., Olsen T.K., Chekenya Enger M., Bjerkvig R.
U-251 revisited: genetic drift and phenotypic consequences of long-term cultures of glioblastoma cells.
Cancer Med. 3:812-824(2014)
PubMed=25960936; DOI=10.4161/21624011.2014.954893; PMCID=PMC4355981
Boegel S., Lower M., Bukur T., Sahin U., Castle J.C.
A catalog of HLA type, HLA expression, and neo-epitope candidates in human cancer cell lines.
OncoImmunology 3:e954893.1-e954893.12(2014)
PubMed=25984343; DOI=10.1038/sdata.2014.35; PMCID=PMC4432652
Cowley G.S., Weir B.A., Vazquez F., Tamayo P., Scott J.A., Rusin S., East-Seletsky A., Ali L.D., Gerath W.F.J., Pantel S.E., Lizotte P.H., Jiang G.-Z., Hsiao J., Tsherniak A., Dwinell E., Aoyama S., Okamoto M., Harrington W., Gelfand E.T., Green T.M., Tomko M.J., Gopal S., Wong T.C., Li H.-B., Howell S., Stransky N., Liefeld T., Jang D., Bistline J., Meyers B.H., Armstrong S.A., Anderson K.C., Stegmaier K., Reich M., Pellman D., Boehm J.S., Mesirov J.P., Golub T.R., Root D.E., Hahn W.C.
Parallel genome-scale loss of function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies.
Sci. Data 1:140035-140035(2014)
PubMed=25894527; DOI=10.1371/journal.pone.0121314; PMCID=PMC4404347
Bausch-Fluck D., Hofmann A., Bock T., Frei A.P., Cerciello F., Jacobs A., Moest H., Omasits U., Gundry R.L., Yoon C., Schiess R., Schmidt A., Mirkowska P., Hartlova A.S., Van Eyk J.E., Bourquin J.-P., Aebersold R., Boheler K.R., Zandstra P.W., Wollscheid B.
A mass spectrometric-derived cell surface protein atlas.
PLoS ONE 10:E0121314-E0121314(2015)
PubMed=27377824; DOI=10.1038/sdata.2016.52; PMCID=PMC4932877
Mestdagh P., Lefever S., Volders P.-J., Derveaux S., Hellemans J., Vandesompele J.
Long non-coding RNA expression profiling in the NCI60 cancer cell line panel using high-throughput RT-qPCR.
Sci. Data 3:160052-160052(2016)
PubMed=27397505; DOI=10.1016/j.cell.2016.06.017; PMCID=PMC4967469
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., Aben N., Goncalves E., Barthorpe S., Lightfoot H., Cokelaer T., Greninger P., van Dyk E., Chang H., de Silva H., Heyn H., Deng X.-M., Egan R.K., Liu Q.-S., Mironenko T., Mitropoulos X., Richardson L., Wang J.-H., Zhang T.-H., Moran S., Sayols S., Soleimani M., Tamborero D., Lopez-Bigas N., Ross-Macdonald P., Esteller M., Gray N.S., Haber D.A., Stratton M.R., Benes C.H., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Garnett M.J.
A landscape of pharmacogenomic interactions in cancer.
Cell 166:740-754(2016)
PubMed=27582061; DOI=10.1126/scitranslmed.aaf6853
Allen M., Bjerke M., Edlund H., Nelander S., Westermark B.
Origin of the U87MG glioma cell line: good news and bad news.
Sci. Transl. Med. 8:354re3.1-354re3.4(2016)
PubMed=27807467; DOI=10.1186/s13100-016-0078-4; PMCID=PMC5087121
Zampella J.G., Rodic N., Yang W.R., Huang C.R.L., Welch J., Gnanakkan V.P., Cornish T.C., Boeke J.D., Burns K.H.
A map of mobile DNA insertions in the NCI-60 human cancer cell panel.
Mob. DNA 7:20.1-20.11(2016)
PubMed=28196595; DOI=10.1016/j.ccell.2017.01.005; PMCID=PMC5501076
Li J., Zhao W., Akbani R., Liu W.-B., Ju Z.-L., Ling S.-Y., Vellano C.P., Roebuck P., Yu Q.-H., Eterovic A.K., Byers L.A., Davies M.A., Deng W.-L., Gopal Y.N.V., Chen G., von Euw E.M., Slamon D.J., Conklin D., Heymach J.V., Gazdar A.F., Minna J.D., Myers J.N., Lu Y.-L., Mills G.B., Liang H.
Characterization of human cancer cell lines by reverse-phase protein arrays.
Cancer Cell 31:225-239(2017)
PubMed=29716531; DOI=10.1186/s12885-018-4394-6; PMCID=PMC5930953
Hegge B., Sjottem E., Mikkola I.
Generation of a PAX6 knockout glioblastoma cell line with changes in cell cycle distribution and sensitivity to oxidative stress.
BMC Cancer 18:496.1-496.19(2018)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747; PMCID=PMC6445675
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=30971826; DOI=10.1038/s41586-019-1103-9
Behan F.M., Iorio F., Picco G., Goncalves E., Beaver C.M., Migliardi G., Santos R., Rao Y., Sassi F., Pinnelli M., Ansari R., Harper S., Jackson D.A., McRae R., Pooley R., Wilkinson P., van der Meer D.J., Dow D., Buser-Doepner C.A., Bertotti A., Trusolino L., Stronach E.A., Saez-Rodriguez J., Yusa K., Garnett M.J.
Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens.
Nature 568:511-516(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3; PMCID=PMC6697103
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. 3rd, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=33385022; DOI=10.1016/j.dib.2020.106643; PMCID=PMC7772531
de Sousa J.F., da Silva P., Serafim R.B., Nociti R.P., Moreira C.G., Silva W.A., Valente V.
RNA sequencing data of different grade astrocytoma cell lines.
Data Brief 34:106643.1-106643.13(2021)
PubMed=35839778; DOI=10.1016/j.ccell.2022.06.010; PMCID=PMC9387775
Goncalves E., Poulos R.C., Cai Z.-X., Barthorpe S., Manda S.S., Lucas N., Beck A., Bucio-Noble D., Dausmann M., Hall C., Hecker M., Koh J., Lightfoot H., Mahboob S., Mali I., Morris J., Richardson L., Seneviratne A.J., Shepherd R., Sykes E., Thomas F., Valentini S., Williams S.G., Wu Y.-X., Xavier D., MacKenzie K.L., Hains P.G., Tully B., Robinson P.J., Zhong Q., Garnett M.J., Reddel R.R.
Pan-cancer proteomic map of 949 human cell lines.
Cancer Cell 40:835-849.e8(2022)
| 产品名称 | 价格 | 指令 |
| Hs683人脑视神经胶质瘤细胞(STR鉴定)[细胞+500ml专培套餐促销] | ¥980.00 | 购物车 》 |
| WERI-Rb-1人视网膜神经胶质瘤细胞 (STR鉴定)[细胞+500ml专培套餐促销] | ¥980.00 | 购物车 》 |
| 支原体检测试剂盒(买二送一) | ¥800.00 | 购物车 》 |
| 支原体清除试剂盒 | ¥600.00 | 购物车 》 |
| 国产转染试剂 | ¥800.00 | 购物车 》 |
| 慢病毒介导基因沉默或过表达 | ¥询价 | 购物车 》 |
| 荧光素酶标记 | ¥询价 | 购物车 》 |
| DMEM高糖基础培养基 | ¥56.00 | 购物车 》 |
| DMEM高糖完全培养基(10%FBS) | ¥350.00 | 购物车 》 |
| 优级胎牛血清 | ¥2580(开学促销) | 购物车 》 |
| 人源细胞STR鉴定服务 | ¥900.00 | 购物车 》 |
 上海中乔新舟生物科技有限公司
上海中乔新舟生物科技有限公司